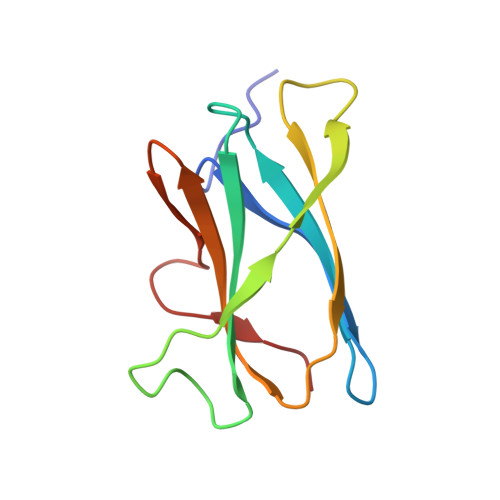
Solution structure of family 21 carbohydrate-binding module from Rhizopus oryzae glucoamylase
Liu, Y.N., Lai, Y.T., Chou, W.I., Chang, M.D.T., Lyu, P.C.(2007) Biochem J 403: 21-30
- PubMed: 17117925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20061312
- Primary Citation Related Structures:
2DJM - PubMed Abstract:
CBMs (carbohydrate-binding modules) function independently to assist carbohydrate-active enzymes. Family 21 CBMs contain approx. 100 amino acid residues, and some members have starchbinding functions or glycogen-binding activities. We report here the first structure of a family 21 CBM from the SBD (starch-binding domain) of Rhizopus oryzae glucoamylase (RoCBM21) determined by NMR spectroscopy. This CBM has a beta-sandwich fold with an immunoglobulin-like structure. Ligand-binding properties of RoCBM21 were analysed by chemical-shift perturbations and automated docking. Structural comparisons with previously reported SBDs revealed two types of topologies, namely type I and type II, with CBM20, CBM25, CBM26 and CBM41 showing type I topology, with CBM21 and CBM34 showing type II topology. According to the chemical-shift perturbations, RoCBM21 contains two ligand-binding sites. Residues in site II are similar to those found in the family 20 CBM from Aspergillus niger glucoamylase (AnCBM20). Site I, however, is embedded in a region with unique sequence motifs only found in some members of CBM21s. Additionally, docking of beta-cyclodextrin and malto-oligosaccharides highlights that side chains of Y83 and W47 (one-letter amino acid code) form the central part of the conserved binding platform in the SBD. The structure of RoCBM21 provides the first direct evidence of the structural features and the basis for protein-carbohydrate recognition from an SBD of CBM21.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, No. 101, Sec. 2, Kuang Fu Rd, Hsinchu, Taiwan 30013.
Organizational Affiliation: