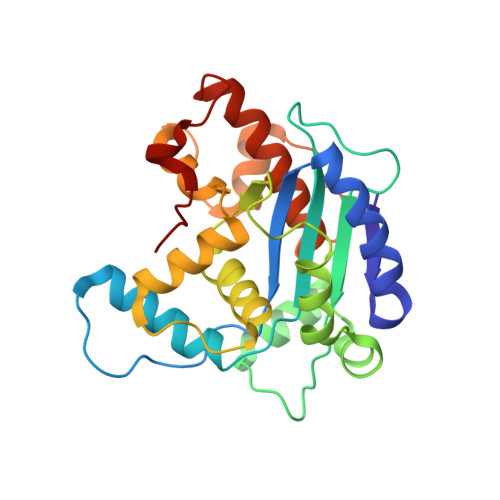
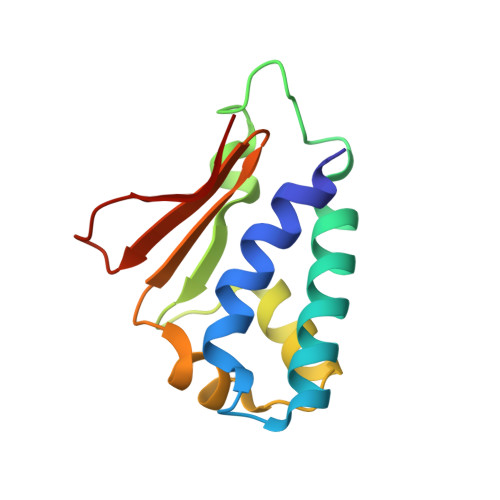
Structural basis for the antiproliferative activity of the Tob-hCaf1 complex.
Horiuchi, M., Takeuchi, K., Noda, N., Muroya, N., Suzuki, T., Nakamura, T., Kawamura-Tsuzuku, J., Takahasi, K., Yamamoto, T., Inagaki, F.(2009) J Biological Chem 284: 13244-13255
- PubMed: 19276069 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M809250200
- Primary Citation Related Structures:
2D5R - PubMed Abstract:
The Tob/BTG family is a group of antiproliferative proteins containing two highly homologous regions, Box A and Box B. These proteins all associate with CCR4-associated factor 1 (Caf1), which belongs to the ribonuclease D (RNase D) family of deadenylases and is a component of the CCR4-Not deadenylase complex. Here we determined the crystal structure of the complex of the N-terminal region of Tob and human Caf1 (hCaf1). Tob exhibited a novel fold, whereas hCaf1 most closely resembled the catalytic domain of yeast Pop2 and human poly(A)-specific ribonuclease. Interestingly, the association of hCaf1 was mediated by both Box A and Box B of Tob. Cell growth assays using both wild-type and mutant proteins revealed that deadenylase activity of Caf1 is not critical but complex formation is crucial to cell growth inhibition. Caf1 tethers Tob to the CCR4-Not deadenylase complex, and thereby Tob gathers several factors at its C-terminal region, such as poly(A)-binding proteins, to exert antiproliferative activity.
- Department of Structural Biology, Graduate School of Pharmaceutical Sciences, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: