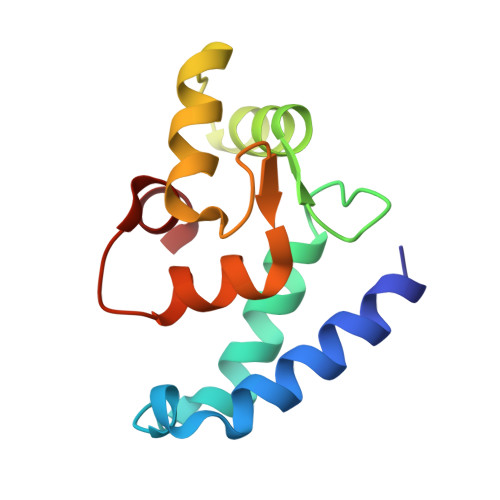
X-ray Structures of the Microglia/Macrophage-specific Protein Iba1 from Human and Mouse Demonstrate Novel Molecular Conformation Change Induced by Calcium binding
Yamada, M., Ohsawa, K., Imai, Y., Kohsaka, S., Kamitori, S.(2006) J Mol Biology 364: 449-457
- PubMed: 17011575
- DOI: https://doi.org/10.1016/j.jmb.2006.09.027
- Primary Citation of Related Structures:
1WY9, 2D58 - PubMed Abstract:
The ionized calcium-binding adaptor molecule 1 (Iba1) with 147 amino acid residues has been identified as a calcium-binding protein, expressed specifically in microglia/macrophages, and is expected to be a key factor in membrane ruffling, which is a typical feature of activated microglia. We have determined the crystal structure of human Iba1 in a Ca(2+)-free form and mouse Iba1 in a Ca(2+)-bound form, to a resolution of 1.9 A and 2.1 A, respectively. X-ray structures of Iba1 revealed a compact, single-domain protein with two EF-hand motifs, showing similarity in overall topology to partial structures of the classical EF-hand proteins troponin C and calmodulin. In mouse Iba1, the second EF-hand contains a bound Ca(2+), but the first EF-hand does not, which is often the case in S100 proteins, suggesting that Iba1 has S100 protein-like EF-hands. The molecular conformational change induced by Ca(2+)-binding of Iba1 is different from that found in the classical EF-hand proteins and/or S100 proteins, which demonstrates that Iba1 has an unique molecular switching mechanism dependent on Ca(2+)-binding, to interact with target molecules.
- Information Technology Center and Faculty of Medicine, Kagawa University, 1750-1 Ikenobe, Miki-cho, Kita-gun, Kagawa 761-0793, Japan.
Organizational Affiliation: