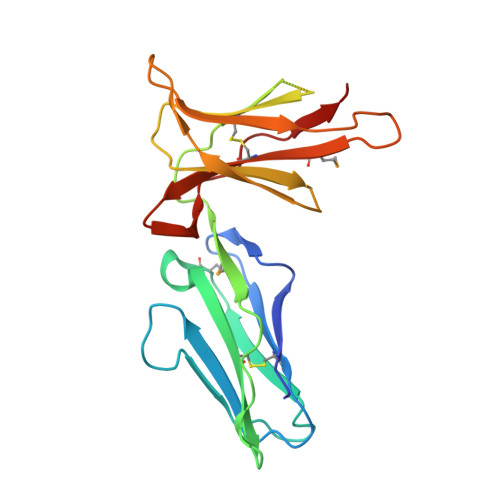
Crystal structure of the human monocyte-activating receptor,
Shiroishi, M., Kajikawa, M., Kuroki, K., Ose, T., Kohda, D., Maenaka, K.(2006) J Biological Chem 281: 19536-19544
- PubMed: 16675463 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M603076200
- Primary Citation Related Structures:
2D3V - PubMed Abstract:
Human leukocyte Ig-like receptor B1 (LILRB1) and B2 (LILRB2) belong to "Group 1" receptors and recognize a broad range of major histocompatibility complex class I molecules (MHCIs). In contrast, "Group 2" receptors show low similarity with LILRB1/B2, and their ligands remain to be identified. To date, the structural and functional characteristics of Group 2 LILRs are poorly understood. Here we report the crystal structure of the extracellular domain of LILRA5, which is an activating Group 2 LILR expressed on monocytes and neutrophils. Unexpectedly, the structure showed large changes in structural conformation and charge distribution in the region corresponding to the MHCI binding site of LILRB1/B2, which are also distinct from killer cell Ig-like receptors and Fc alpha receptors. These changes probably confer the structural hindrance for the MHCI binding, and their key amino acid substitutions are well conserved in Group 2 LILRs. Consistently, the surface plasmon resonance and flow cytometric analyses demonstrated that LILRA5 exhibited no affinities to all tested MHCIs. These results raised the possibility that LILRA5 as well as Group 2 LILRs do not play a role in any MHCI recognition but could possibly bind to non-MHCI ligand(s) on the target cells to provide a novel immune regulation mechanism.
- Division of Structural Biology, Medical Institute of Bioregulation, Kyushu University, 3-1-1 Maidashi, Higashi-ku, Fukuoka 812-8582, Japan.
Organizational Affiliation: