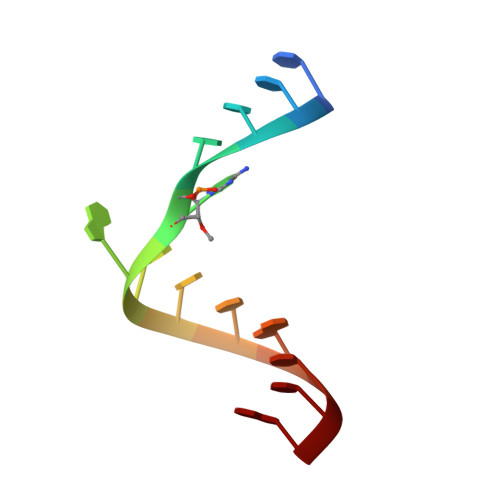
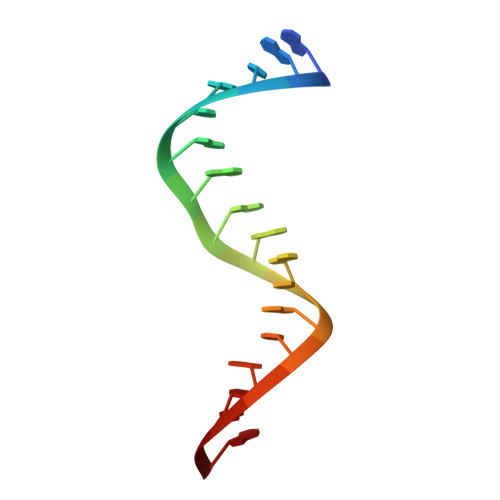
Conformational heterogeneity at position U37 of an all-RNA hairpin ribozyme with implications for metal binding and the catalytic structure of the S-turn
Alam, S., Grum-Tokars, V., Krucinska, J., Kundracik, M.L., Wedekind, J.E.(2005) Biochemistry 44: 14396-14408
- PubMed: 16262240 Search on PubMed
- DOI: https://doi.org/10.1021/bi051550i
- Primary Citation Related Structures:
1X9C, 1X9K, 2D2K, 2D2L - PubMed Abstract:
The hairpin ribozyme is an RNA enzyme that performs site-specific phosphodiester bond cleavage between nucleotides A-1 and G+1 within its cognate substrate. Previous functional studies revealed that the minimal hairpin ribozyme exhibited "gain-of-function" cleavage properties resulting from U39C or U39 to propyl linker (C3) modifications. Furthermore, each "mutant" displayed different magnesium-dependence in its activity. To investigate the molecular basis for these gain-of-function variants, crystal structures of minimal, junctionless hairpin ribozymes were solved in native (U39), and mutant U39C and U39(C3) forms. The results revealed an overall molecular architecture comprising two docked internal loop domains folded into a wishbone shape, whose tertiary interface forms a sequestered active site. All three minimal hairpin ribozymes bound Co(NH(3))(6)(3+) at G21/A40, the E-loop/S-turn boundary. The native structure also showed that U37 of the S-turn adopts both sequestered and exposed conformations that differ by a maximum displacement of 13 A. In the sequestered form, the U37 base packs against G36, and its 2'-hydroxyl group forms a water mediated hydrogen bond to O4' of G+1. These interactions were not observed in previous four-way-junction hairpin ribozyme structures due to crystal contacts with the U1A splicing protein. Interestingly, the U39C and U39(C3) mutations shifted the equilibrium conformation of U37 into the sequestered form through formation of new hydrogen bonds in the S-turn, proximal to the essential nucleotide A38. A comparison of all three new structures has implications for the catalytically relevant conformation of the S-turn and suggests a rationale for the distinctive metal dependence of each mutant.
- Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry, Rochester, New York 14642, USA.
Organizational Affiliation: