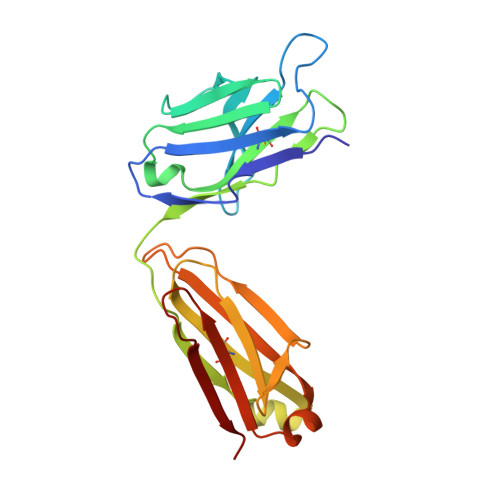
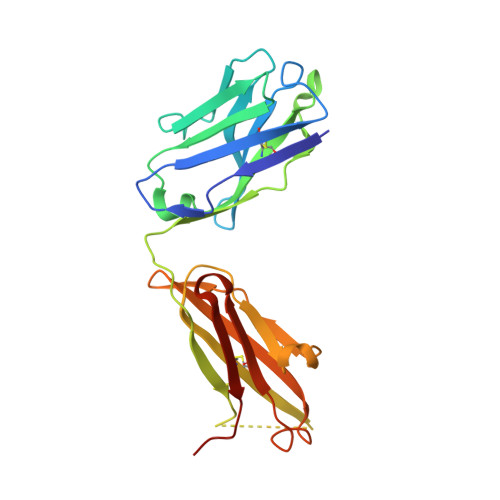
Crystallographic analysis of the NNA7 Fab and proposal for the mode of human blood-group recognition.
Xie, K., Song, S.C., Spitalnik, S.L., Wedekind, J.E.(2005) Acta Crystallogr D Biol Crystallogr 61: 1386-1394
- PubMed: 16204891 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905023851
- Primary Citation Related Structures:
2D03 - PubMed Abstract:
The NNA7 Fab antibody fragment recognizes the human N-type blood-group antigen comprised of the N-terminal glycopeptide of glycophorin A (GPA). A mutant form of this Fab fragment, NNA7-G91S, exhibits markedly reduced antigen binding. To provide insight into how these Fab fragments recognize this glycopeptide antigen, the crystal structures of NNA7 and NNA7-G91S were solved and refined to 1.83 and 1.97 A resolution, respectively. Both molecules are composed of the same heavy (H) chain Fd fragment, but each contains a slightly different light (L) chain owing to the G91S substitution. Specifically, the G91S mutation pushes the backbone of the neighboring H chain away from complementarity-determining region 3 (CDR3) of the L-chain variable region, allowing an additional glycerol cryoprotectant molecule to enter the antigen-combining site near the putative location of O-linked glycosylation. Each Fab fragment also possesses a well defined 2-(N-morpholino)ethanesulfonic acid (MES) molecule trapped in its antigen-combining site, as well as a crystallographic symmetry-related molecule comprising an amino-acid sequence that is virtually identical to the N-terminus of GPA. The MES molecule interacts with the H-chain CDR in a manner reminiscent of antibody-carbohydrate complexes. These results suggest a model for recognition of the glycopeptide antigen that accounts for the deleterious effect of the G91S substitution.
- Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry, Rochester, NY 14642, USA.
Organizational Affiliation: