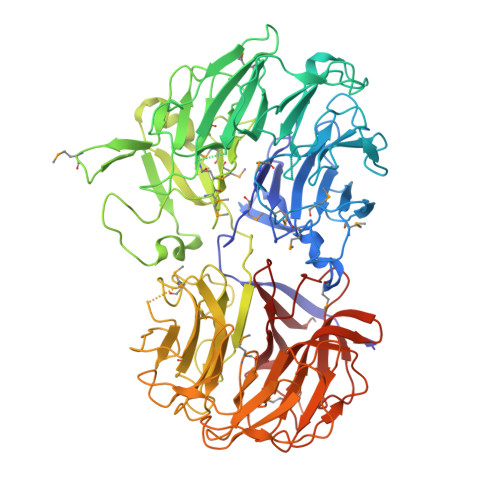
Crystal Structures of Clostridium Thermocellum Xyloglucanase, Xgh74A, Reveal the Structural Basis for Xyloglucan Recognition and Degradation.
Martinez-Fleites, C., Guerreiro, C.I., Baumann, M.J., Taylor, E.J., Prates, J.A.M., Ferreira, L.M.A., Fontes, C.M.G.A., Brumer, H., Davies, G.J.(2006) J Biological Chem 281: 24922
- PubMed: 16772298 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M603583200
- Primary Citation Related Structures:
2CN2, 2CN3 - PubMed Abstract:
The enzymatic degradation of the plant cell wall is central both to the natural carbon cycle and, increasingly, to environmentally friendly routes to biomass conversion, including the production of biofuels. The plant cell wall is a complex composite of cellulose microfibrils embedded in diverse polysaccharides collectively termed hemicelluloses. Xyloglucan is one such polysaccharide whose hydrolysis is catalyzed by diverse xyloglucanases. Here we present the structure of the Clostridium thermocellum xyloglucanase Xgh74A in both apo and ligand-complexed forms. The structures, in combination with mutagenesis data on the catalytic residues and the kinetics and specificity of xyloglucan hydrolysis reveal a complex subsite specificity accommodating seventeen monosaccharide moieties of the multibranched substrate in an open substrate binding terrain.
- York Structural Biology Laboratory, Department of Chemitry, University of York, York YO10 5YW, United Kingdom.
Organizational Affiliation: