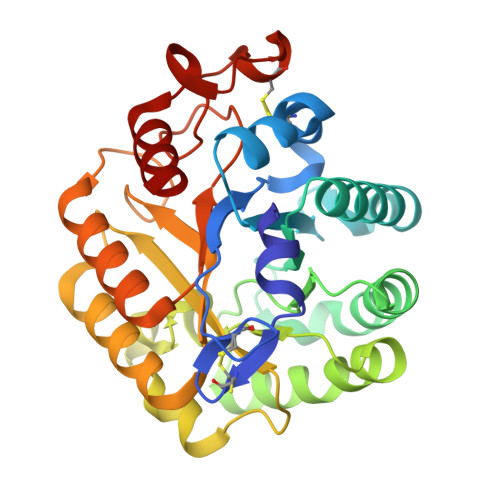
The Crystal Structure of the Catalytic Domain of Thermobifida Fusca Endoglucanase Cel5A in Complex with Cellotetraose
Berglund, G.I., Gualfetti, P.J., Requadt, C., Gross, L.S., Bergfors, T., Shaw, A., Saldajeno, M., Mitchinson, C., Sandgren, M.To be published.