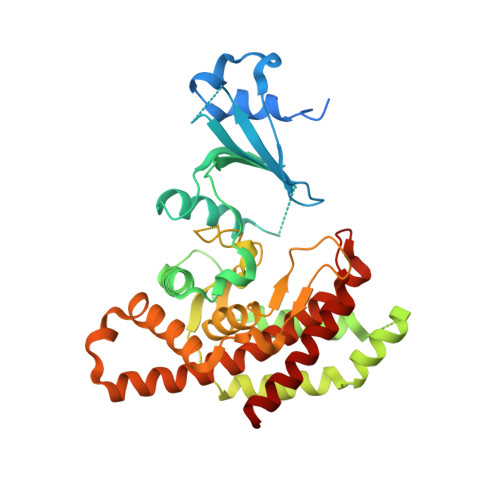
Elucidation of Human Choline Kinase Crystal Structures in Complex with the Products Adp or Phosphocholine.
Malito, E., Sekulic, N., Too, W.C., Konrad, M., Lavie, A.(2006) J Mol Biology 364: 136
- PubMed: 17007874 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.08.084
- Primary Citation Related Structures:
2CKO, 2CKP, 2CKQ - PubMed Abstract:
Choline kinase, responsible for the phosphorylation of choline to phosphocholine as the first step of the CDP-choline pathway for the biosynthesis of phosphatidylcholine, has been recognized as a new target for anticancer therapy. Crystal structures of human choline kinase in its apo, ADP and phosphocholine-bound complexes, respectively, reveal the molecular details of the substrate binding sites. ATP binds in a cavity where residues from both the N and C-terminal lobes contribute to form a cleft, while the choline-binding site constitutes a deep hydrophobic groove in the C-terminal domain with a rim composed of negatively charged residues. Upon binding of choline, the enzyme undergoes conformational changes independently affecting the N-terminal domain and the ATP-binding loop. From this structural analysis and comparison with other kinases, and from mutagenesis data on the homologous Caenorhabditis elegans choline kinase, a model of the ternary ADP.phosphocholine complex was built that reveals the molecular basis for the phosphoryl transfer activity of this enzyme.
- Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, Chicago, IL 60607, USA.
Organizational Affiliation: