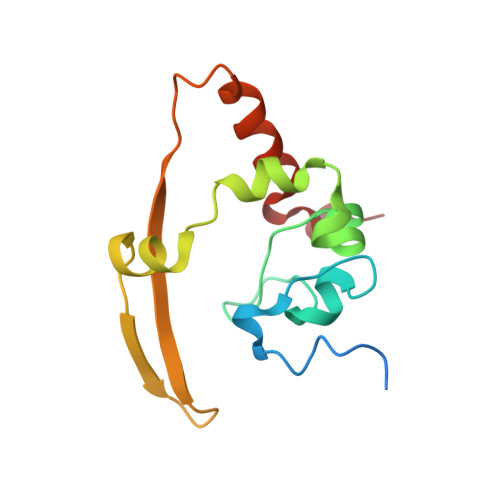
Structure of the Sars Coronavirus Nucleocapsid Protein RNA-Binding Dimerization Domain Suggests a Mechanism for Helical Packaging of Viral RNA.
Chen, C.-Y., Chang, C.K., Chang, Y.W., Sue, S.C., Bai, H.I., Riang, L., Hsiao, C.-D., Huang, T.H.(2007) J Mol Biology 368: 1075
- PubMed: 17379242
- DOI: https://doi.org/10.1016/j.jmb.2007.02.069
- Primary Citation Related Structures:
2CJR - PubMed Abstract:
Coronavirus nucleocapsid proteins are basic proteins that encapsulate viral genomic RNA to form part of the virus structure. The nucleocapsid protein of SARS-CoV is highly antigenic and associated with several host-cell interactions. Our previous studies using nuclear magnetic resonance revealed the domain organization of the SARS-CoV nucleocapsid protein. RNA has been shown to bind to the N-terminal domain (NTD), although recently the C-terminal half of the protein has also been implicated in RNA binding. Here, we report that the C-terminal domain (CTD), spanning residues 248-365 (NP248-365), had stronger nucleic acid-binding activity than the NTD. To determine the molecular basis of this activity, we have also solved the crystal structure of the NP248-365 region. Residues 248-280 form a positively charged groove similar to that found in the infectious bronchitis virus (IBV) nucleocapsid protein. Furthermore, the positively charged surface area is larger in the SARS-CoV construct than in the IBV. Interactions between residues 248-280 and the rest of the molecule also stabilize the formation of an octamer in the asymmetric unit. Packing of the octamers in the crystal forms two parallel, basic helical grooves, which may be oligonucleotide attachment sites, and suggests a mechanism for helical RNA packaging in the virus.
- Institute of Molecular Biology, Academia Sinica, Taipei 115, Taiwan, ROC.
Organizational Affiliation: