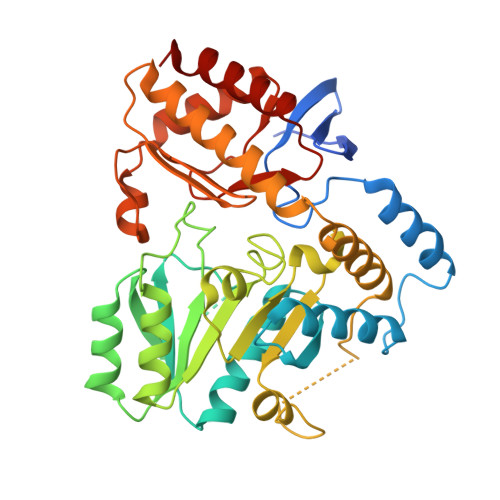
Evolutionary Relationship between Initial Enzymes of Tetrapyrrole Biosynthesis
Schulze, J.O., Schubert, W.-D., Moser, J., Jahn, D., Heinz, D.W.(2006) J Mol Biology 358: 1212
- PubMed: 16564539 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.02.064
- Primary Citation Related Structures:
2CFB - PubMed Abstract:
Glutamate-1-semialdehyde 2,1-aminomutase (GSAM) is the second enzyme in the C(5) pathway of tetrapyrrole biosynthesis found in most bacteria, in archaea and in plants. It catalyzes the transamination of glutamate-1-semialdehyde to 5-aminolevulinic acid (ALA) in a pyridoxal 5'-phosphate (PLP)-dependent manner. We present the crystal structure of GSAM from the thermophilic cyanobacterium Thermosynechococcus elongatus (GSAM(Tel)) in its PLP-bound form at 2.85A resolution. GSAM(Tel) is a symmetric homodimer, whereas GSAM from Synechococcus (GSAM(Syn)) has been described as asymmetric. The symmetry of GSAM(Tel) thus challenges the previously proposed negative cooperativity between monomers of this enzyme. Furthermore, GSAM(Tel) reveals an extensive flexible region at the interface of the proposed complex of GSAM with glutamyl-tRNA reductase (GluTR), the preceding enzyme in tetrapyrrole biosynthesis. Compared to GSAM(Syn), the monomers of GSAM(Tel) are rotated away from each other along the dimerization interface by 10 degrees . The associated flexibility of GSAM may be essential for complex formation with GluTR to occur. Unexpectedly, we find that GSAM is structurally related to 5-aminolevulinate synthase (ALAS), the ALA-producing enzyme in the Shemin pathway of alpha-proteobacteria and non-plant eukaryotes. This structural relationship applies also to the corresponding subfamilies of PLP-dependent enzymes. We thus propose that the CoA-subfamily (including ALAS) and the aminotransferase subfamily II (including GSAM) are evolutionarily closely related and that ALAS may thus have evolved from GSAM.
- Division of Structural Biology, German Research Centre for Biotechnology (GBF), Mascheroder Weg 1, D-38124 Braunschweig, Germany.
Organizational Affiliation: