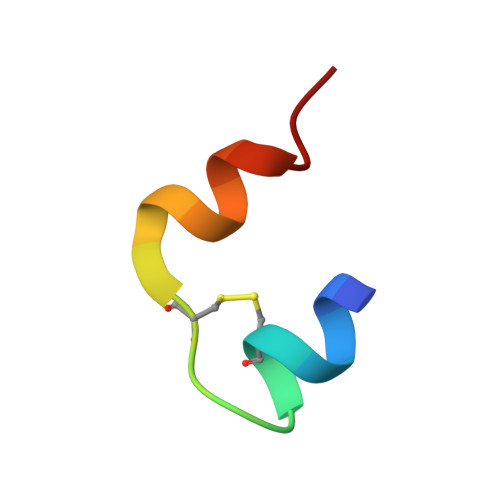
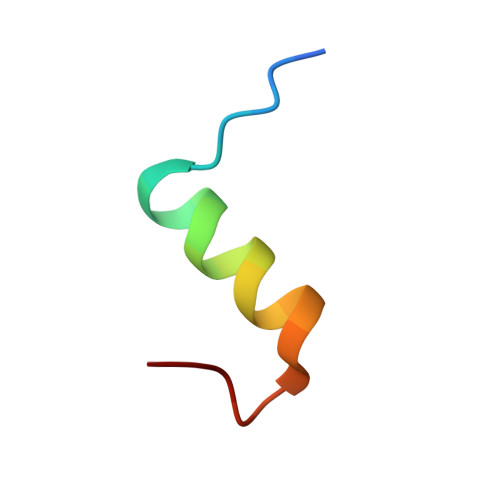
I222 Crystal Form of Despentapeptide (B26-B30) Insulin Provides New Insights Into the Properties of Monomeric Insulin.
Whittingham, J.L., Youshang, Z., Zakova, L., Dodson, E.J., Turkenburg, J.P., Brange, J., Dodson, G.G.(2006) Acta Crystallogr D Biol Crystallogr 62: 505
- PubMed: 16627943 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444906006871
- Primary Citation Related Structures:
2CEU - PubMed Abstract:
Despentapeptide (des-B26-B30) insulin (DPI), an active modified insulin, has been crystallized in the presence of 20% acetic acid pH 2. A crystal structure analysis to 1.8 A spacing (space group I222) revealed that the DPI molecule, which is unable to make beta-strand interactions for physiological dimer formation and is apparently monomeric in solution, formed an alternative lattice-generated dimer. The formation of this dimer involved interactions between surfaces which included the B9-B19 alpha-helices (usually buried by the dimer-dimer contacts within the native hexamer). The two crystallographically independent molecules within the dimer were essentially identical and were similar in conformation to T-state insulin as seen in the T(6) insulin hexamer. An unusual feature of each molecule in the dimer was the presence of two independent conformations at the B-chain C-terminus (residues B20-B25). Both conformations were different from that of native insulin, involving a 3.5 A displacement of the B20-B23 beta-turn and a repositioning of residue PheB25 such that it made close van der Waals contact with the main body of the molecule, appearing to stabilize the B-chain C-terminus.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5YW, England.
Organizational Affiliation: