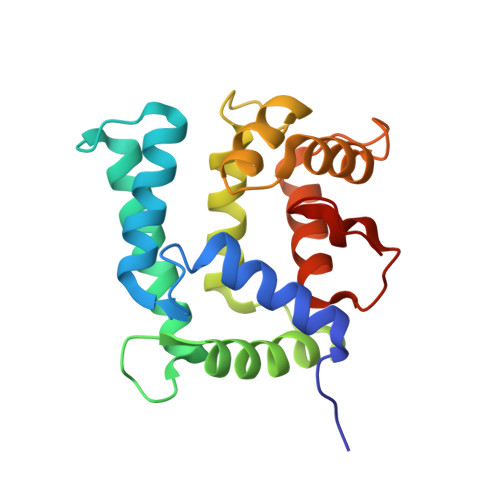
Structure of the Neuronal Protein Calexcitin Suggests a Mode of Interaction in Signalling Pathways of Learning and Memory.
Erskine, P.T., Beaven, G.D.E., Hagan, R., Findlow, I.S., Werner, J.M., Wood, S.P., Vernon, J., Giese, K.P., Fox, G., Cooper, J.B.(2006) J Mol Biology 357: 1536
- PubMed: 16497326 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.01.083
- Primary Citation Related Structures:
2CCM - PubMed Abstract:
The three-dimensional structure of the neuronal calcium-sensor protein calexcitin from Loligo pealei has been determined by X-ray analysis at a resolution of 1.8A. Calexcitin is up-regulated following Pavlovian conditioning and has been shown to regulate potassium channels and the ryanodine receptor. Thus, calexcitin is implicated in neuronal excitation and plasticity. The overall structure is predominantly helical and compact with a pronounced hydrophobic core between the N and C-terminal domains of the molecule. The structure consists of four EF-hand motifs although only the first three EF hands are involved in binding calcium ions; the C-terminal EF-hand lacks the amino acids required for calcium binding. The overall structure is quite similar to that of the sarcoplasmic calcium-binding protein from Amphioxus although the sequence identity is very low at 31%. The structure shows that the two amino acids of calexcitin phosphorylated by protein kinase C are close to the domain interface in three dimensions and thus phosphorylation is likely to regulate the opening of the domains that is probably required for binding to target proteins. There is evidence that calexcitin is a GTPase and the residues, which have been implicated by mutagenesis in its GTPase activity, are in a short but highly conserved region of 3(10) helix close to the C terminus. This helix resides in a large loop that is partly sandwiched between the N and C-terminal domains suggesting that GTP binding may also require or may cause domain opening. The structure possesses a pronounced electropositive crevice in the vicinity of the 3(10) helix, that might provide an initial docking site for the triphosphate group of GTP. These findings elucidate a number of the reported functions of calexcitin with implications for neuronal signalling.
- School of Biological Sciences, University of Southampton, Bassett Crescent East, Southampton SO16 7PX, UK.
Organizational Affiliation: