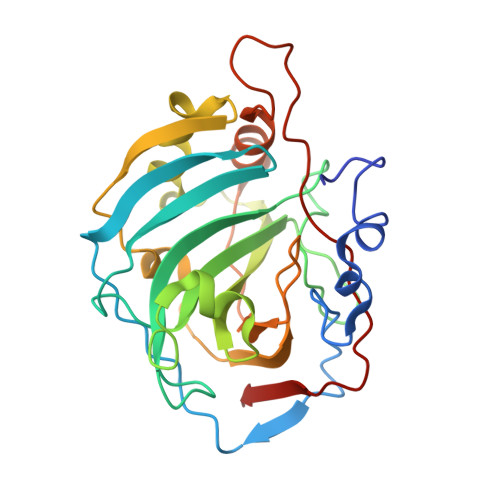
Structure, refinement, and function of carbonic anhydrase isozymes: refinement of human carbonic anhydrase I
Kannan, K.K., Ramanadham, M., Jones, T.A.(1984) Ann N Y Acad Sci 429: 49-60
- PubMed: 6430186 Search on PubMed
- DOI: https://doi.org/10.1111/j.1749-6632.1984.tb12314.x
- Primary Citation Related Structures:
2CAB - PubMed Abstract:
The structure of human erythrocyte carbonic anhydrase I has been refined to a final R value of 19% to 2-A resolution by a combination of least squares refinement and model fitting in a three-dimensional graphics display. About 300 solvent atoms have been located bound to the protein molecule. An interesting hydrogen bond network involving Zn2+, the liganded solvent, side chain groups of Thr-199, Glu-106, Thr-7, and His-64 through two solvent molecules have been found that may be important for the catalytic mechanism of the carbonic anhydrase.