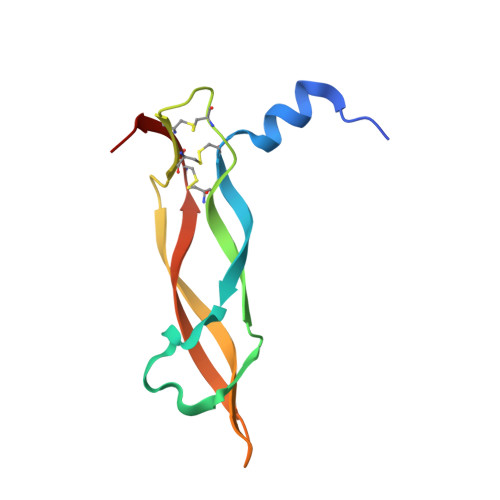
Crystal Structure of Human Vascular Endothelial Growth Factor-B: Identification of Amino Acids Important for Receptor Binding
Iyer, S., Scotney, P.D., Nash, A.D., Acharya, K.R.(2006) J Mol Biology 359: 76
- PubMed: 16616187 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.03.002
- Primary Citation Related Structures:
2C7W - PubMed Abstract:
The development of blood vessels (angiogenesis) is critical throughout embryogenesis and in some normal postnatal physiological processes. Pathological angiogenesis has a pivotal role in sustaining tumour growth and chronic inflammation. Vascular endothelial growth factor-B (VEGF-B) is a member of the VEGF family of growth factors that regulate blood vessel and lymphatic angiogenesis. VEGF-B is closely related to VEGF-A and placenta growth factor (PlGF), but unlike VEGF-A, which binds to two receptor tyrosine kinases VEGFR-1 (Flt-1) and VEGFR-2 (Flk-1/KDR), VEGF-B and PlGF bind to VEGFR-1 and not VEGFR-2. There is growing evidence of a role for VEGF-B in physiological and pathological blood vessel angiogenesis. VEGF-B may provide novel therapeutic strategies for the treatment of vascular disease and be a potential therapeutic target in aberrant vessel formation. To help understand at the molecular level the differential receptor binding profile of the VEGF family of growth factors we have determined the crystal structure of human VEGF-B(10-108) at 2.48 Angstroms resolution. The overall structure is very similar to that of the previously determined cysteine-knot motif growth factors: VEGF-A, PlGF and platelet-derived growth factor-B (PDGF-B). We also present a predicted model for the association of VEGF-B with the second domain of its receptor, VEGFR-1. Based on this interaction and the present structural data of the native protein, we have identified several putative residues that could play an important role in receptor recognition and specificity.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, UK.
Organizational Affiliation: