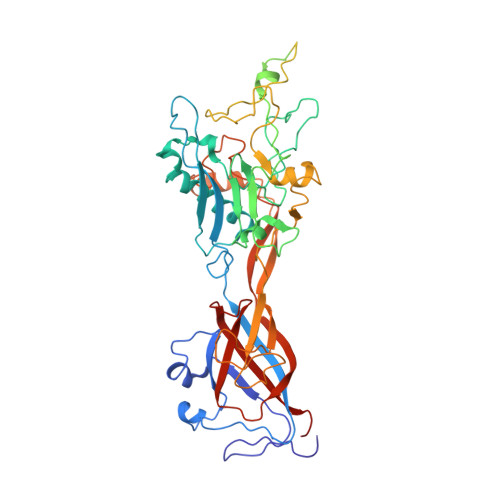
Structural and Biochemical Characterization of a Human Adenovirus 2/12 Penton Base Chimera.
Zubieta, C., Blanchoin, L., Cusack, S.(2006) FEBS J 273: 4336
- PubMed: 16939624 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05430.x
- Primary Citation Related Structures:
2C6S - PubMed Abstract:
The vertex of the adenoviral capsid is formed by the penton, a complex of two proteins, the pentameric penton base and the trimeric fiber protein. The penton contains all necessary components for viral attachment and entry into the host cell. After initial attachment via the head domain of the fiber protein, the penton base interacts with cellular integrins through an Arg-Gly-Asp (RGD) motif located in a hypervariable surface loop, triggering virus internalization. In order to investigate the structural and functional role of this region, we replaced the hypervariable loop of serotype 2 with the corresponding, but much shorter, loop of serotype 12 and compared it to the wild type. Here, we report the 3.6 A crystal structure of a human adenovirus 2/12 penton base chimera crystallized as a dodecamer. The structure is generally similar to human adenovirus 2 penton base, with the main differences localized to the fiber protein-binding site. Fluorescence anisotropy assays using a trimeric fiber protein mimetic called the minifiber and wild-type human adenovirus 2 and chimeric penton base demonstrate that fiber protein binding is independent of the hypervariable loop, with a K(d) for fiber binding estimated in the 1-2 microm range. Interestingly, competition assays using labeled and unlabeled minifiber demonstrated virtually irreversible binding to the penton base, which we ascribe to a conformational change, on the basis of comparisons of all available penton base structures.
- European Molecular Biology Laboratory, Grenoble Outstation, France. czubieta@slac.stanford.edu
Organizational Affiliation: