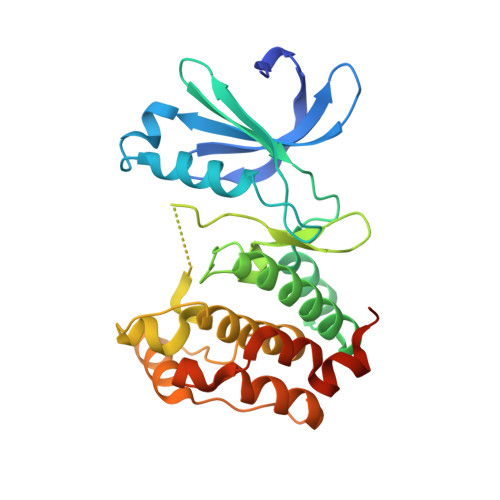
Sar and Inhibitor Complex Structure Determination of a Novel Class of Potent and Specific Aurora Kinase Inhibitors.
Heron, N.M., Anderson, M., Blowers, D.P., Breed, J., Eden, J.M., Green, S., Hill, G.B., Johnson, T., Jung, F.H., Mcmiken, H.H.J., Mortlock, A.A., Pannifer, A.D., Pauptit, R.A., Pink, J., Roberts, N.J., Rowsell, S.(2006) Bioorg Med Chem Lett 16: 1320
- PubMed: 16337122 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2005.11.053
- Primary Citation Related Structures:
2C6D, 2C6E - PubMed Abstract:
A novel series of 5-aminopyrimidinyl quinazolines has been developed from anilino-quinazoline 1, which was identified in a high throughput screen for Aurora A. Introduction of the pyrimidine ring and optimisation of the substituents both on this ring and at the C7 position of the quinazoline led to the discovery of compounds that are highly specific Aurora kinase inhibitors. Co-crystallisation of one of these inhibitors with a fragment of Aurora A shows the importance of the benzamido group in achieving selectivity.
- AstraZeneca, Mereside, Alderley Park, Macclesfield, Cheshire SK10 4TG,UK. Nicola.Heron@astrazeneca.com
Organizational Affiliation: