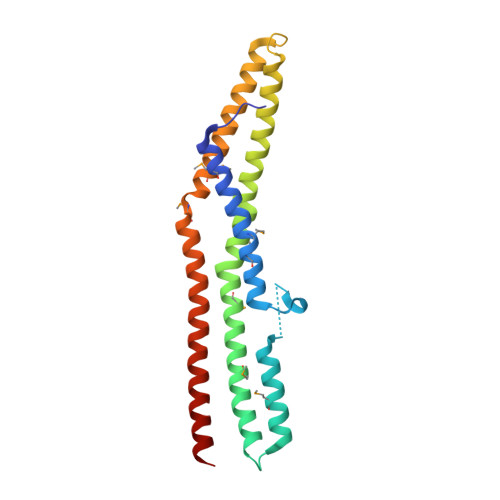
Mechanism of Endophilin N-Bar Domain-Mediated Membrane Curvature.
Gallop, J.L., Jao, C.C., Kent, H.M., Butler, P.J., Evans, P.R., Langen, R., Mcmahon, H.T.(2006) EMBO J 25: 2898
- PubMed: 16763559 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601174
- Primary Citation Related Structures:
2C08 - PubMed Abstract:
Endophilin-A1 is a BAR domain-containing protein enriched at synapses and is implicated in synaptic vesicle endocytosis. It binds to dynamin and synaptojanin via a C-terminal SH3 domain. We examine the mechanism by which the BAR domain and an N-terminal amphipathic helix, which folds upon membrane binding, work as a functional unit (the N-BAR domain) to promote dimerisation and membrane curvature generation. By electron paramagnetic resonance spectroscopy, we show that this amphipathic helix is peripherally bound in the plane of the membrane, with the midpoint of insertion aligned with the phosphate level of headgroups. This places the helix in an optimal position to effect membrane curvature generation. We solved the crystal structure of rat endophilin-A1 BAR domain and examined a distinctive insert protruding from the membrane interaction face. This insert is predicted to form an additional amphipathic helix and is important for curvature generation. Its presence defines an endophilin/nadrin subclass of BAR domains. We propose that N-BAR domains function as low-affinity dimers regulating binding partner recruitment to areas of high membrane curvature.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: