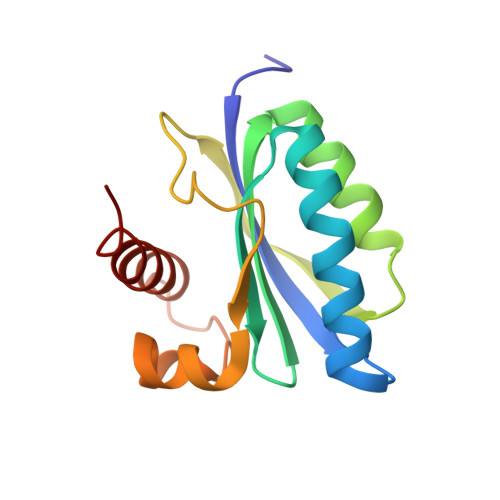
Structure of a Bacterial Bluf Photoreceptor: Insights Into Blue Light-Mediated Signal Transduction.
Jung, A., Domratcheva, T., Tarutina, M., Wu, Q., Ko, W.H., Shoeman, R.L., Gomelsky, M., Gardner, K.H., Schlichting, I.(2005) Proc Natl Acad Sci U S A 102: 12350
- PubMed: 16107542 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0500722102
- Primary Citation Related Structures:
2BYC - PubMed Abstract:
Light is an essential environmental factor, and many species have evolved the capability to respond to it. Blue light is perceived through three flavin-containing photoreceptor families: cryptochromes, light-oxygen-voltage, and BLUF (sensor of blue light using flavin adenine dinucleotide, FAD) domain proteins. BLUF domains are present in various proteins from Bacteria and lower Eukarya. They are fully modular and can relay signals to structurally and functionally diverse output units, most of which are implicated in nucleotide metabolism. We present the high resolution crystal structure of the dark resting state of BlrB, a short BLUF domain-containing protein from Rhodobacter sphaeroides. The structure reveals a previously uncharacterized FAD-binding fold. Along with other lines of evidence, it suggests mechanistic aspects for the photocycle that is characterized by a red-shifted absorbance of the flavin. The isoalloxazine ring of FAD binds in a cleft between two helices, whereas the adenine ring points into the solvent. We propose that the adenine ring serves as a hook mediating the interaction with its effector/output domain. The structure suggests a unique photochemical signaling switch in which the absorption of light induces a structural change in the rim surrounding the hook, thereby changing the protein interface between BLUF and the output domain.
- Max Planck Institute for Medical Research, Department of Biomolecular Mechanisms, Jahnstrasse 29, 69120 Heidelberg, Germany.
Organizational Affiliation: