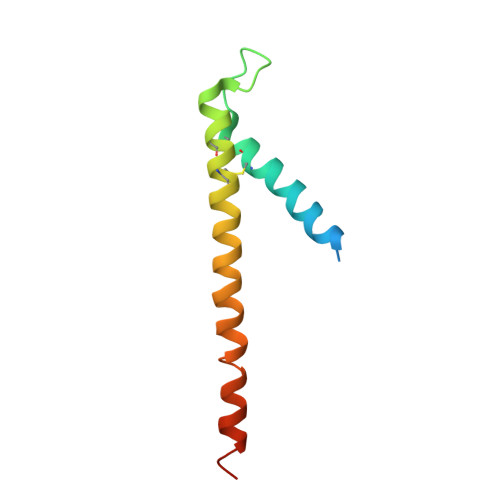
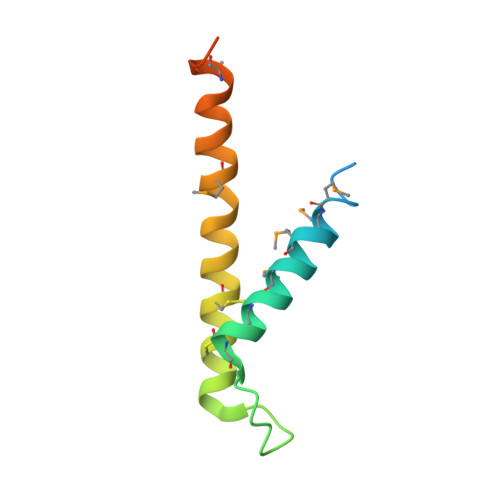
Crystal Structure of the Mitochondrial Chaperone Tim910 Reveals a Six-Bladed Alpha-Propeller.
Webb, C.T., Gorman, M.A., Lazarou, M., Ryan, M.T., Gulbis, J.M.(2006) Mol Cell 21: 123
- PubMed: 16387659 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.11.010
- Primary Citation Related Structures:
2BSK - PubMed Abstract:
Import of proteins into mitochondria occurs by coordinated actions of preprotein translocases in the outer and inner membranes. Tim9 and Tim10 are translocase components of the intermembrane space, related to deafness-dystonia peptide 1 (DDP1). They coassemble into a hexamer, TIM9.10, which captures and chaperones precursors of inner membrane metabolite carriers as they exit the TOM channel in the outer membrane. The crystal structure of TIM9.10 reveals a previously undescribed alpha-propeller topology in which helical "blades" radiate from a narrow central pore lined with polar residues. The propeller blades are reminiscent of "tentacles" in chaperones Skp and prefoldin. In each TIM9.10 subunit, a signature "twin CX3C" motif forms two intramolecular disulfides. There is no obvious binding pocket for precursors, which we suggest employ the chaperone-like tentacles of TIM9.10 as surrogate lipid contacts. The first reported crystal structure of a mitochondrial translocase assembly provides insights into selectivity and regulation of precursor import.
- Structural Biology Division, The Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, Victoria 3050, Australia.
Organizational Affiliation: