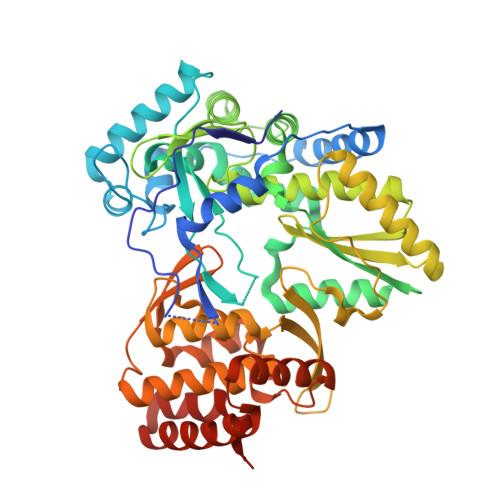
Interdomain Communication in Hepatitis C Virus Polymerase Abolished by Small-Molecule Inhibitors Bound to a Novel Allosteric Site
Di Marco, S., Volpari, C., Tomei, L., Altamura, S., Harper, S., Narjes, F., Koch, U., Rowley, M., De Francesco, R., Migliaccio, G., Carfi, A.(2005) J Biological Chem 280: 29765
- PubMed: 15955819 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M505423200
- Primary Citation Related Structures:
2BRK, 2BRL - PubMed Abstract:
The hepatitis C virus (HCV) polymerase is required for replication of the viral genome and is a key target for therapeutic intervention against HCV. We have determined the crystal structures of the HCV polymerase complexed with two indole-based allosteric inhibitors at 2.3- and 2.4-Angstroms resolution. The structures show that these inhibitors bind to a site on the surface of the thumb domain. A cyclohexyl and phenyl ring substituents, bridged by an indole moiety, fill two closely spaced pockets, whereas a carboxylate substituent forms a salt bridge with an exposed arginine side chain. Interestingly, in the apoenzyme, the inhibitor binding site is occupied by a small alpha-helix at the tip of the N-terminal loop that connects the fingers and thumb domains. Thus, these molecules inhibit the enzyme by preventing formation of intramolecular contacts between these two domains and consequently precluding their coordinated movements during RNA synthesis. Our structures identify a novel mechanism by which a new class of allosteric inhibitors inhibits the HCV polymerase and open the way to the development of novel antiviral agents against this clinically relevant human pathogen.
- Istituto di Ricerche di Biologia Molecolare P. Angeletti, Pomezia (Rome), Italy.
Organizational Affiliation: