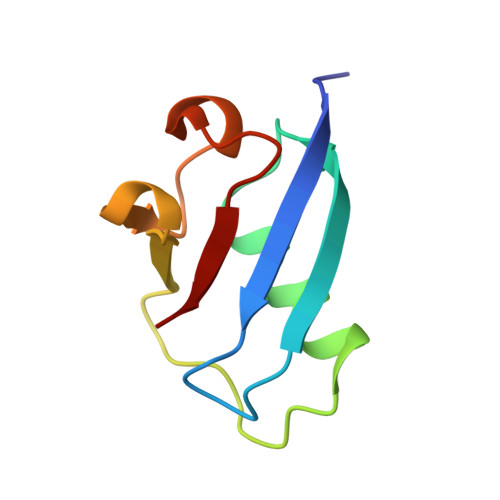
Crystal Structure of Ubiquitin-Like Protein Yukd from Bacillus Subtilis
van den Ent, F., Lowe, J.(2005) FEBS Lett 579: 3837
- PubMed: 15978580 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2005.06.002
- Primary Citation Related Structures:
2BPS - PubMed Abstract:
The YukD protein in Bacillus subtilis was identified in a hidden Markov model (HMM) search as being related in sequence to ubiquitin. By solving the crystal structure we show that YukD adopts a fold that is most closely related to ubiquitin, yet has the shortest C-terminal tail of all known ubiquitin-like proteins. The endogenous gene of yukD in B. subtilis was disrupted without an obvious phenotypic effect and an inducible copy encoding a C-Myc and His-tagged version of the protein was introduced at the ectopic locus amyE. Conjugation assays performed both in vitro and in vivo indicate that YukD lacks the capacity for covalent bond formation with other proteins.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK. fent@mrc-lmb.cam.ac.uk
Organizational Affiliation: