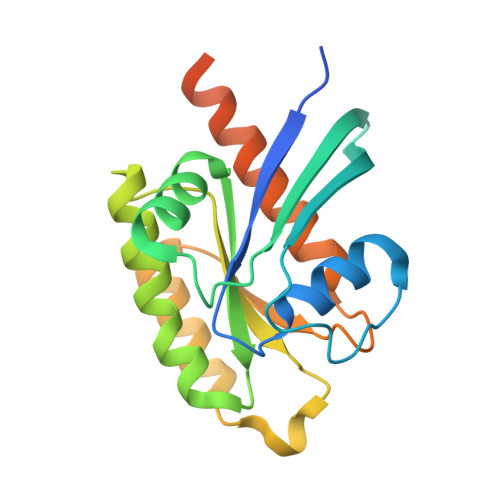
Molecular Recognition of an Adp-Ribosylating Clostridium Botulinum C3 Exoenzyme by Rala Gtpase
Holbourn, K.P., Sutton, J.M., Evans, H.R., Shone, C.C., Acharya, K.R.(2005) Proc Natl Acad Sci U S A 102: 5357
- PubMed: 15809419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0501525102
- Primary Citation Related Structures:
2BOV - PubMed Abstract:
C3 exoenzymes (members of the ADP-ribosyltranferase family) are produced by Clostridium botulinum (C3bot1 and -2), Clostridium limosum (C3lim), Bacillus cereus (C3cer), and Staphylococcus aureus (C3stau1-3). These exoenzymes lack a translocation domain but are known to specifically inactivate Rho GTPases in host target cells. Here, we report the crystal structure of C3bot1 in complex with RalA (a GTPase of the Ras subfamily) and GDP at a resolution of 2.66 A. RalA is not ADP-ribosylated by C3 exoenzymes but inhibits ADP-ribosylation of RhoA by C3bot1, C3lim, and C3cer to different extents. The structure provides an insight into the molecular interactions between C3bot1 and RalA involving the catalytic ADP-ribosylating turn-turn (ARTT) loop from C3bot1 and helix alpha4 and strand beta6 (which are not part of the GDP-binding pocket) from RalA. The structure also suggests a molecular explanation for the different levels of C3-exoenzyme inhibition by RalA and why RhoA does not bind C3bot1 in this manner.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: