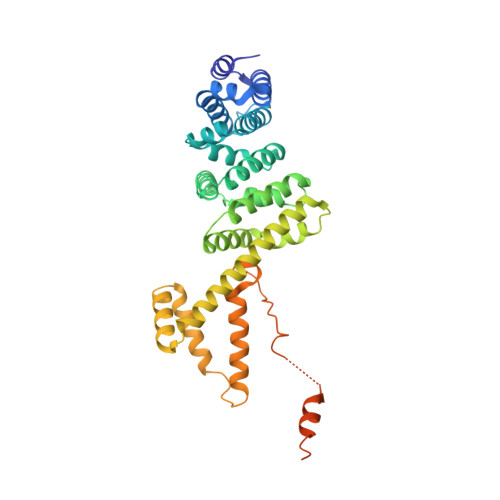
Structural Basis of Rho Gtpase-Mediated Activation of the Formin Mdia1
Otomo, T., Otomo, C., Tomchick, D.R., Machius, M., Rosen, M.K.(2005) Mol Cell 18: 273
- PubMed: 15866170 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.04.002
- Primary Citation Related Structures:
2BNX - PubMed Abstract:
Diaphanous-related formins (DRFs) regulate dynamics of unbranched actin filaments during cell contraction and cytokinesis. DRFs are autoinhibited through intramolecular binding of a Diaphanous autoinhibitory domain (DAD) to a conserved N-terminal regulatory element. Autoinhibition is relieved through binding of the GTPase RhoA to the N-terminal element. We report the crystal structure of the dimeric regulatory domain of the DRF, mDia1. Dimerization is mediated by an intertwined six-helix bundle, from which extend two Diaphanous inhibitory domains (DIDs) composed of five armadillo repeats. NMR and biochemical mapping indicate the RhoA and DAD binding sites on the DID partially overlap, explaining activation of mDia1 by the GTPase. RhoA binding also requires an additional structurally independent segment adjacent to the DID. This regulatory construction, involving a GTPase binding site spanning a flexibly tethered arm and the inhibitory module, is observed in many autoinhibited effectors of Ras superfamily GTPases, suggesting evolutionary pressure for this design.
- Department of Biochemistry, University of Texas Southwestern Medical Center at Dallas, 5323 Harry Hines Boulevard, Dallas, Texas 75390, USA.
Organizational Affiliation: