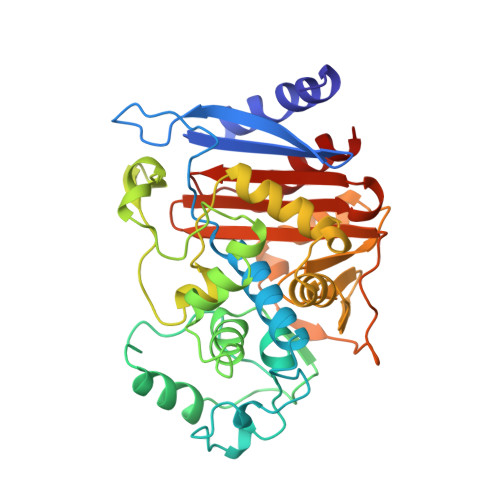
Three-dimensional structure of AmpC beta-lactamase from Escherichia coli bound to a transition-state analogue: possible implications for the oxyanion hypothesis and for inhibitor design.
Usher, K.C., Blaszczak, L.C., Weston, G.S., Shoichet, B.K., Remington, S.J.(1998) Biochemistry 37: 16082-16092
- PubMed: 9819201 Search on PubMed
- DOI: https://doi.org/10.1021/bi981210f
- Primary Citation Related Structures:
2BLS, 3BLS - PubMed Abstract:
The structures of AmpC beta-lactamase from Escherichia coli, alone and in complex with a transition-state analogue, have been determined by X-ray crystallography. The native enzyme was determined to 2.0 A resolution, and the structure with the transition-state analogue m-aminophenylboronic acid was determined to 2.3 A resolution. The structure of AmpC from E. coli resembles those previously determined for the class C enzymes from Enterobacter cloacae and Citrobacter freundii. The transition-state analogue, m-aminophenylboronic acid, makes several interactions with AmpC that were unexpected. Perhaps most surprisingly, the putative "oxyanion" of the boronic acid forms what appears to be a hydrogen bond with the backbone carbonyl oxygen of Ala318, suggesting that this atom is protonated. Although this interaction has not previously been discussed, a carbonyl oxygen contact with the putative oxyanion or ligand carbonyl oxygen appears in most complexes involving a beta-lactam recognizing enzyme. These observations may suggest that the high-energy intermediate for amide hydrolysis by beta-lactamases and related enzymes involves a hydroxyl and not an oxyanion, although the oxyanion form certainly cannot be discounted. The involvement of the main-chain carbonyl in ligand and transition-state recognition is a distinguishing feature between serine beta-lactamases and serine proteases, to which they are often compared. AmpC may use the interaction between the carbonyl of Ala318 and the carbonyl of the acylated enzyme to destabilize the ground-state intermediate, this destabilization energy might be relieved in the transition state by a hydroxyl hydrogen bond. The structure of the m-aminophenylboronic acid adduct also suggests several ways to improve the affinity of this class of inhibitor and points to the existence of several unusual binding-site-like features in the region of the AmpC catalytic site.
- Institute of Molecular Biology, University of Oregon, Eugene 97403, USA.
Organizational Affiliation: