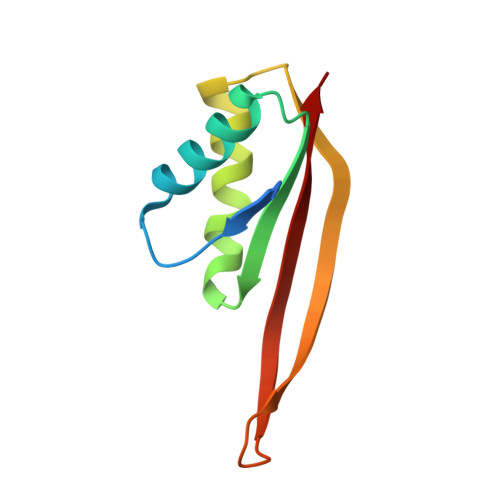
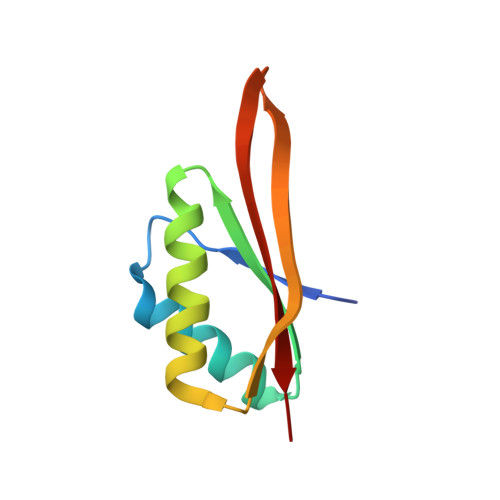
Obligate Heterodimerization of the Archaeal Alba2 Protein with Alba1 Provides a Mechanism for Control of DNA Packaging.
Jelinska, C., Conroy, M.J., Craven, C.J., Hounslow, A.M., Bullough, P.A., Waltho, J.P., Taylor, G.L., White, M.F.(2005) Structure 13: 963
- PubMed: 16004869 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.04.016
- Primary Citation Related Structures:
2BKY - PubMed Abstract:
Organisms growing at elevated temperatures face a particular challenge to maintain the integrity of their genetic material. All thermophilic and hyperthermophilic archaea encode one or more copies of the Alba (Sac10b) gene. Alba is an abundant, dimeric, highly basic protein that binds cooperatively and at high density to DNA. Sulfolobus solfataricus encodes a second copy of the Alba gene, and the Alba2 protein is expressed at approximately 5% of the level of Alba1. We demonstrate by NMR, ITC, and crystallography that Alba2 exists exclusively as a heterodimer with Alba1 at physiological concentrations and that heterodimerization exerts a clear effect upon the DNA packaging, as observed by EM, potentially by changing the interface between adjacent Alba dimers in DNA complexes. A functional role for Alba2 in modulation of higher order chromatin structure and DNA condensation is suggested.
- Centre for Biomolecular Science, University of Saint Andrews, North Haugh, Saint Andrews, Fife KY16 9ST, United Kingdom.
Organizational Affiliation: