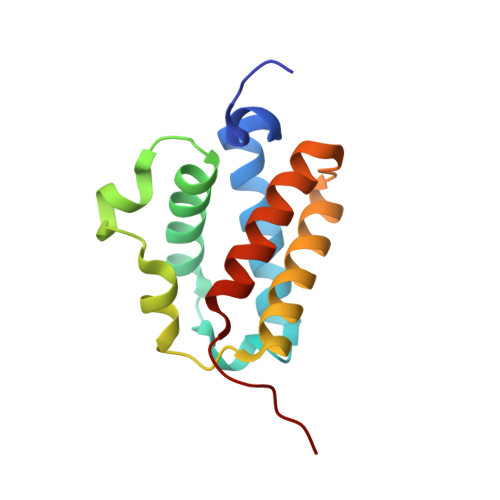
Crystal Structure and Ligand Binding Properties of the Truncated Hemoglobin from Geobacillus Stearothermophilus
Ilari, A., Kjelgaard, P., von Wachenfeldt, C., Catacchio, B., Chiancone, E., Boffi, A.(2007) Arch Biochem Biophys 457: 85
- PubMed: 17126283 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2006.09.033
- Primary Citation Related Structures:
2BKM - PubMed Abstract:
A novel truncated hemoglobin has been identified in the thermophilic bacterium Geobacillus stearothermophilus (Gs-trHb). The protein has been expressed in Escherichia coli, the 3D crystal structure (at 1.5 Angstroms resolution) and the ligand binding properties have been determined. The distal heme pocket displays an array of hydrogen bonding donors to the iron-bound ligands, including Tyr-B10 on one side of the heme pocket and Trp-G8 indole nitrogen on the opposite side. At variance with the highly similar Bacillus subtilis hemoglobin, Gs-trHb is dimeric both in the crystal and in solution and displays several unique structural properties. In the crystal cell, the iron-bound ligand is not homogeneously distributed within each distal site such that oxygen and an acetate anion can be resolved with relative occupancies of 50% each. Accordingly, equilibrium titrations of the oxygenated derivative in solution with acetate anion yield a partially saturated ferric acetate adduct. Moreover, the asymmetric unit contains two subunits and sedimentation velocity ultracentrifugation data confirm that the protein is dimeric.
- Department of Biochemical Sciences and CNR Institute of Molecular Biology and Pathology, University of Rome La Sapienza, P le Aldo Moro 5, 00185 Rome, Italy.
Organizational Affiliation: