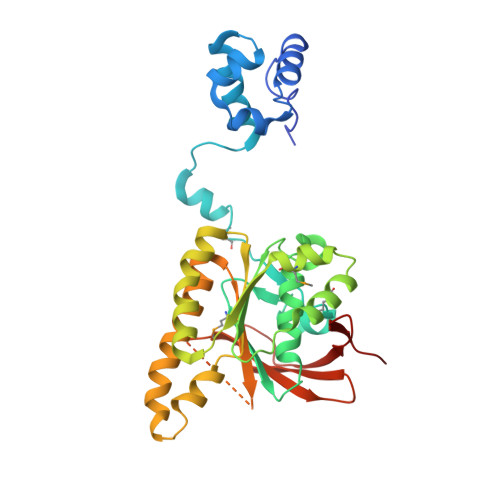
Conformational Flexibility Revealed by the Crystal Structure of a Crenarchaeal Rada
Ariza, A., Richard, D.L., White, M.F., Bond, C.S.(2005) Nucleic Acids Res 33: 1465
- PubMed: 15755748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki288
- Primary Citation Related Structures:
2BKE - PubMed Abstract:
Homologous recombinational repair is an essential mechanism for repair of double-strand breaks in DNA. Recombinases of the RecA-fold family play a crucial role in this process, forming filaments that utilize ATP to mediate their interactions with single- and double-stranded DNA. The recombinase molecules present in the archaea (RadA) and eukaryota (Rad51) are more closely related to each other than to their bacterial counterpart (RecA) and, as a result, RadA makes a suitable model for the eukaryotic system. The crystal structure of Sulfolobus solfataricus RadA has been solved to a resolution of 3.2 A in the absence of nucleotide analogues or DNA, revealing a narrow filamentous assembly with three molecules per helical turn. As observed in other RecA-family recombinases, each RadA molecule in the filament is linked to its neighbour via interactions of a short beta-strand with the neighbouring ATPase domain. However, despite apparent flexibility between domains, comparison with other structures indicates conservation of a number of key interactions that introduce rigidity to the system, allowing allosteric control of the filament by interaction with ATP. Additional analysis reveals that the interaction specificity of the five human Rad51 paralogues can be predicted using a simple model based on the RadA structure.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee Dow St, Dundee, DD1 5EH, UK.
Organizational Affiliation: