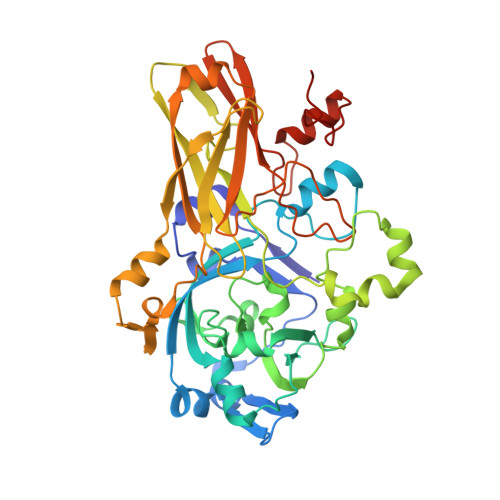
Structural Basis of Eukaryotic Nitrate Reduction: Crystal Structures of the Nitrate Reductase Active Site
Fischer, K., Barbier, G., Hecht, H.-J., Mendel, R.R., Campbell, W.H., Schwarz, G.(2005) Plant Cell 17: 1167
- PubMed: 15772287 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1105/tpc.104.029694
- Primary Citation Related Structures:
2BIH, 2BII - PubMed Abstract:
Nitrate assimilation in autotrophs provides most of the reduced nitrogen on earth. In eukaryotes, reduction of nitrate to nitrite is catalyzed by the molybdenum-containing NAD(P)H:nitrate reductase (NR; EC 1.7.1.1-3). In addition to the molybdenum center, NR contains iron-heme and flavin adenine dinucleotide as redox cofactors involved in an internal electron transport chain from NAD(P)H to nitrate. Recombinant, catalytically active Pichia angusta nitrate-reducing, molybdenum-containing fragment (NR-Mo) was expressed in P. pastoris and purified. Crystal structures for NR-Mo were determined at 1.7 and 2.6 angstroms. These structures revealed a unique slot for binding nitrate in the active site and identified key Arg and Trp residues potentially involved in nitrate binding. Dimeric NR-Mo is similar in overall structure to sulfite oxidases, with significant differences in the active site. Sulfate bound in the active site caused conformational changes, as compared with the unbound enzyme. Four ordered water molecules located in close proximity to Mo define a nitrate binding site, a penta-coordinated reaction intermediate, and product release. Because yeast NAD(P)H:NR is representative of the family of eukaryotic NR, we propose a general mechanism for nitrate reduction catalysis.
- Institute of Plant Biology, Technical University Braunschweig, D-38106 Braunschweig, Germany.
Organizational Affiliation: