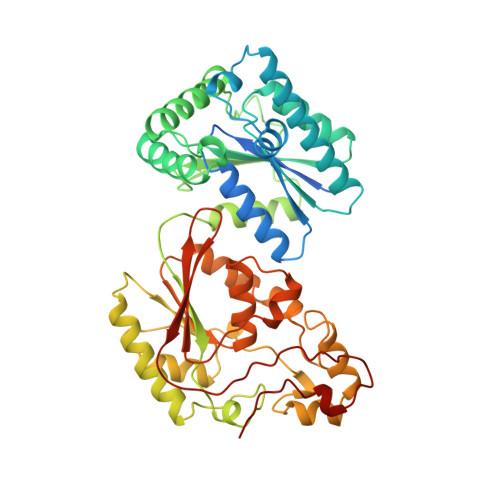
Crystal structure of the H256A mutant of rat testis fructose-6-phosphate,2-kinase/fructose-2,6-bisphosphatase. Fructose 6-phosphate in the active site leads to mechanisms for both mutant and wild type bisphosphatase activities.
Yuen, M.H., Mizuguchi, H., Lee, Y.H., Cook, P.F., Uyeda, K., Hasemann, C.A.(1999) J Biological Chem 274: 2176-2184
- PubMed: 9890980 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.4.2176
- Primary Citation Related Structures:
2BIF - PubMed Abstract:
Fructose-6-phosphate,2-kinase/fructose-2,6-bisphosphatase (Fru-6-P, 2-kinase/Fru-2,6-Pase) is a bifunctional enzyme, catalyzing the interconversion of beta-D-fructose- 6-phosphate (Fru-6-P) and fructose-2,6-bisphosphate (Fru-2,6-P2) at distinct active sites. A mutant rat testis isozyme with an alanine replacement for the catalytic histidine (H256A) in the Fru-2,6-Pase domain retains 17% of the wild type activity (Mizuguchi, H., Cook, P. F., Tai, C-H., Hasemann, C. A., and Uyeda, K. (1998) J. Biol. Chem. 274, 2166-2175). We have solved the crystal structure of H256A to a resolution of 2. 4 A by molecular replacement. Clear electron density for Fru-6-P is found at the Fru-2,6-Pase active site, revealing the important interactions in substrate/product binding. A superposition of the H256A structure with the RT2K-Wo structure reveals no significant reorganization of the active site resulting from the binding of Fru-6-P or the H256A mutation. Using this superposition, we have built a view of the Fru-2,6-P2-bound enzyme and identify the residues responsible for catalysis. This analysis yields distinct catalytic mechanisms for the wild type and mutant proteins. The wild type mechanism would lead to an inefficient transfer of a proton to the leaving group Fru-6-P, which is consistent with a view of this event being rate-limiting, explaining the extremely slow turnover (0. 032 s-1) of the Fru-2,6-Pase in all Fru-6-P,2-kinase/Fru-2,6-Pase isozymes.
- Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas 75235, USA.
Organizational Affiliation: