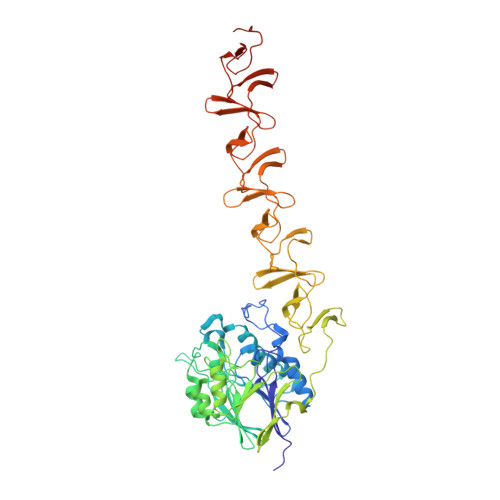
Insights Into Pneumococcal Pathogenesis from Crystal Structure of the Modular Teichoic Acid Phosphorylcholine Esterase Pce
Hermoso, J.A., Lagartera, L., Gonzalez, A., Stelter, M., Garcia, P., Martinez-Ripoll, M., Garcia, J.L., Menendez, M.(2005) Nat Struct Mol Biol 12: 533
- PubMed: 15895092 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb940
- Primary Citation Related Structures:
2BIB - PubMed Abstract:
Phosphorylcholine, a specific component of the pneumococcal cell wall, is crucial in pathogenesis. It directly binds to the human platelet-activating factor (PAF) receptor and acts as a docking station for the family of surface-located choline-binding proteins (CBP). The first structure of a complete pneumococcal CBP, Pce (or CbpE), has been solved in complex with the reaction product and choline analogs. Pce has a novel modular structure, with a globular N-terminal module containing a binuclear Zn(2+) catalytic center, and an elongated choline-binding module. Residues involved in substrate binding and catalysis are described and modular configuration of the active center accounts for in vivo features of teichoic acid hydrolysis. The hydrolysis of PAF by Pce and its regulatory role in phosphorylcholine decoration of the bacterial surface provide new insights into the critical function of Pce in pneumococcal adherence and invasiveness.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Química-Física Rocasolano, CSIC, Serrano 119, 28006 Madrid, Spain. xjuan@iqfr.csic.es
Organizational Affiliation: