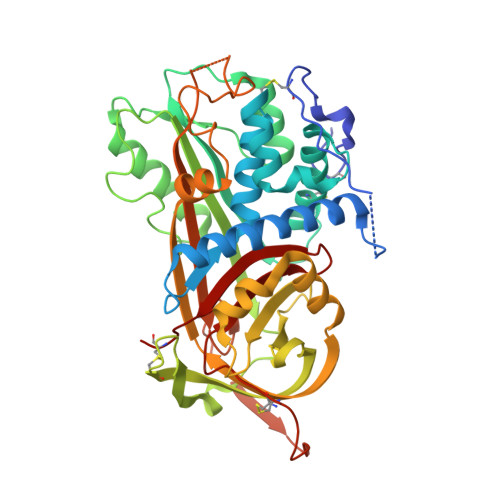
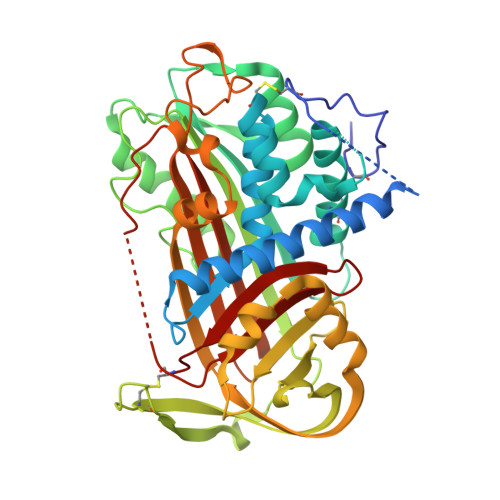
Crystal structure of monomeric native antithrombin reveals a novel reactive center loop conformation.
Johnson, D.J., Langdown, J., Li, W., Luis, S.A., Baglin, T.P., Huntington, J.A.(2006) J Biological Chem 281: 35478-35486
- PubMed: 16973611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M607204200
- Primary Citation Related Structures:
1T1F, 2B5T, 2BEH - PubMed Abstract:
The poor inhibitory activity of circulating antithrombin (AT) is critical to the formation of blood clots at sites of vascular damage. AT becomes an efficient inhibitor of the coagulation proteases only after binding to a specific heparin pentasaccharide, which alters the conformation of the reactive center loop (RCL). The molecular basis of this activation event lies at the heart of the regulation of hemostasis and accounts for the anticoagulant properties of the low molecular weight heparins. Although several structures of AT have been solved, the conformation of the RCL in native AT remains unknown because of the obligate crystal contact between the RCL of native AT and its latent counterpart. Here we report the crystallographic structure of a variant of AT in its monomeric native state. The RCL shifted approximately 20 A, and a salt bridge was observed between the P1 residue (Arg-393) and Glu-237. This contact explains the effect of mutations at the P1 position on the affinity of AT for heparin and also the properties of AT-Truro (E237K). The relevance of the observed conformation was verified through mutagenesis studies and by solving structures of the same variant in different crystal forms. We conclude that the poor inhibitory activity of the circulating form of AT is partially conferred by intramolecular contacts that restrain the RCL, orient the P1 residue away from attacking proteases, and additionally block the exosite utilized in protease recognition.
- Department of Haematology, Division of Structural Medicine, Thrombosis Research Unit, Cambridge Institute for Medical Research, University of Cambridge, Wellcome Trust/MRC Building, Hills Road, Cambridge CB2 2XY, United Kingdom.
Organizational Affiliation: