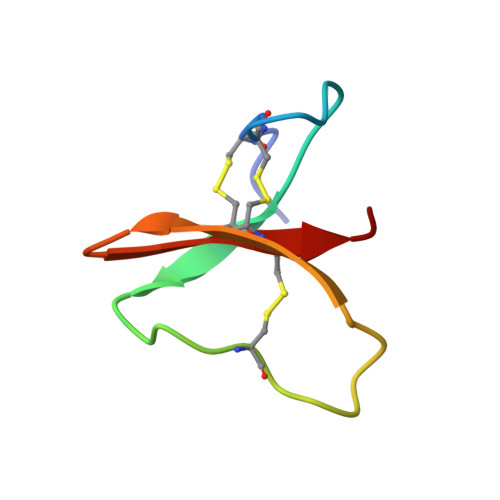
Determination of the three-dimensional solution structure of the antihypertensive and antiviral protein BDS-I from the sea anemone Anemonia sulcata: a study using nuclear magnetic resonance and hybrid distance geometry-dynamical simulated annealing.
Driscoll, P.C., Gronenborn, A.M., Beress, L., Clore, G.M.(1989) Biochemistry 28: 2188-2198
- PubMed: 2566326 Search on PubMed
- DOI: https://doi.org/10.1021/bi00431a033
- Primary Citation Related Structures:
1BDS, 2BDS - PubMed Abstract:
The three-dimensional solution structure of the antihypertensive and antiviral protein BDS-I from the sea anemone Anemonia sulcata has been determined on the basis of 489 interproton and 24 hydrogen-bonding distance restraints supplemented by 23 phi backbone and 21 chi 1 side-chain torsion angle restraints derived from nuclear magnetic resonance (NMR) measurements. A total of 42 structures is calculated by a hybrid metric matrix distance geometry-dynamical simulated annealing approach. Both the backbone and side-chain atom positions are well defined. The average atomic rms difference between the 42 individual SA structures and the mean structure obtained by averaging their coordinates is 0.67 +/- 0.12 A for the backbone atoms and 0.90 +/- 0.17 A for all atoms. The core of the protein is formed by a triple-stranded antiparallel beta-sheet composed of residues 14-16 (strand 1), 30-34 (strand 2), and 37-41 (strand 3) with an additional mini-antiparallel beta-sheet at the N-terminus (residues 6-9). The first and second strands of the triple-stranded antiparallel beta-sheet are connected by a long exposed loop (residues 17-30). A number of side-chain interactions are discussed in light of the structure.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, Bethesda, Maryland 20892.
Organizational Affiliation: