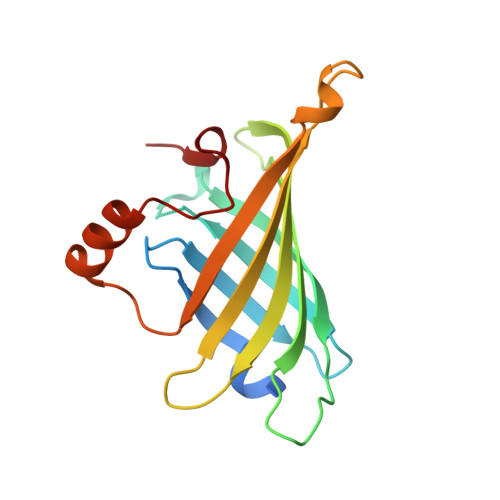
Crystallographic Analysis of a Full-length Streptavidin with Its C-terminal Polypeptide Bound in the Biotin Binding Site.
Le Trong, I., Humbert, N., Ward, T.R., Stenkamp, R.E.(2006) J Mol Biology 356: 738-745
- PubMed: 16384581 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.086
- Primary Citation Related Structures:
2BC3 - PubMed Abstract:
The structure of a full-length streptavidin has been determined at 1.7 A resolution and shows that the 20 residue extension at the C terminus forms a well-ordered polypeptide loop on the surface of the tetramer. Residues 150-153 of the extension are bound to the ligand-binding site, possibly competing with exogenous ligands. The binding mode of these residues is compared with that of biotin and peptidic ligands. The observed structure helps to rationalize the observations that full-length mature streptavidin binds biotinylated macromolecules with reduced affinity.
- Departments of Biological Structure and Biochemistry and the Biomolecular Structure Center, University of Washington, Box 357420, Seattle, WA 98195-7420, USA.
Organizational Affiliation: