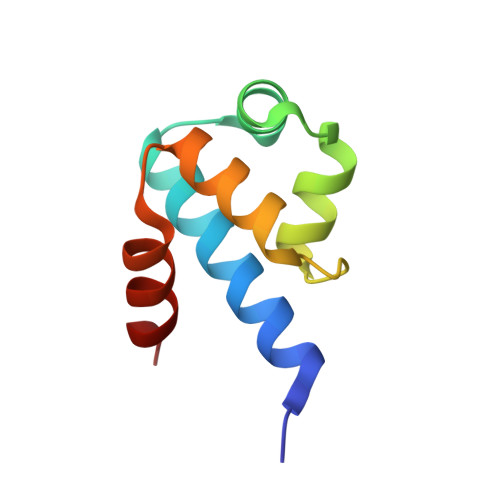
Crystal Structure of the Restriction-Modification System Control Element C.BclI and Mapping of Its Binding Site.
Sawaya, M.R., Zhu, Z., Mersha, F., Chan, S.H., Dabur, R., Xu, S.Y., Balendiran, G.K.(2005) Structure 13: 1837-1847
- PubMed: 16338412 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.08.017
- Primary Citation Related Structures:
2B5A - PubMed Abstract:
Protection from DNA invasion is afforded by restriction-modification systems in many bacteria. The efficiency of protection depends crucially on the relative expression levels of restriction versus methytransferase genes. This regulation is provided by a controller protein, named C protein. Studies of the Bcll system in E. coli suggest that C.Bcll functions as a negative regulator for M.Bcll expression, implying that it plays a role in defense against foreign DNA during virus infection. C.Bcll binds (Kd = 14.3 nM) to a 2-fold symmetric C box DNA sequence that overlaps with the putative -35 promoter region upstream of the bcllM and bcllC genes. The C.Bcll fold comprises five alpha helices: two helices form a helix-turn-helix motif, and the remaining three helices form the extensive dimer interface. The C.Bcll-DNA model proposed suggests that DNA bending might play an important role in gene regulation, and that Glu27 and Asp31 in C.Bcll might function critically in the regulation.
- UCLA-DOE Laboratory of Structural Biology and Molecular Medicine, 205 Boyer Hall, Box 951570, Los Angeles, California 90095, USA.
Organizational Affiliation: