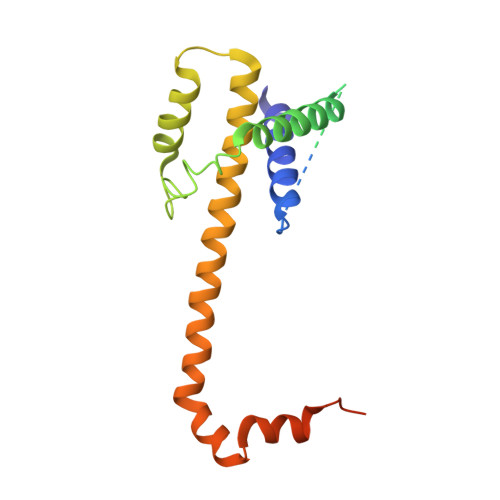
BCL-X(L) Dimerization by Three-dimensional Domain Swapping.
O'Neill, J.W., Manion, M.K., Maguire, B., Hockenbery, D.M.(2006) J Mol Biology 356: 367-381
- PubMed: 16368107 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.032
- Primary Citation Related Structures:
2B48 - PubMed Abstract:
Dimeric interactions among anti- and pro-apoptotic members of the BCL-2 protein family are dynamically regulated and intimately involved in survival and death functions. We report the structure of a BCL-X(L) homodimers a 3D-domain swapped dimer (3DDS). The X-ray crystal structure demonstrates the mutual exchange of carboxy-terminal regions including BH2 (Bcl-2 homology 2) between monomer subunits, with the hinge region occurring at the hairpin turn between the fifth and sixth alpha helices. Both BH3 peptide-binding hydrophobic grooves are unoccupied in the 3DDS dimer and available for BH3 peptide binding, as confirmed by sedimentation velocity analysis. BCL-X(L) 3DDS dimers have increased pore-forming activity compared to monomers, suggesting that 3DDS dimers may act as intermediates in membrane pore formation. Chemical crosslinking studies of Cys-substituted BCL-X(L) proteins demonstrate that 3DDS dimers form in synthetic lipid vesicles.
- Divisions of Clinical Research and Human Biology, Fred Hutchinson Cancer Research Center, Seattle, WA, USA.
Organizational Affiliation: