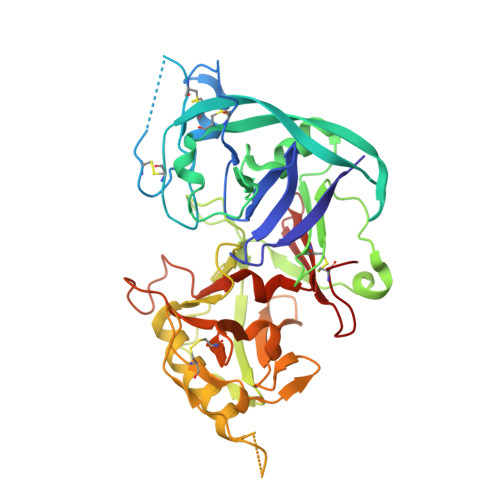
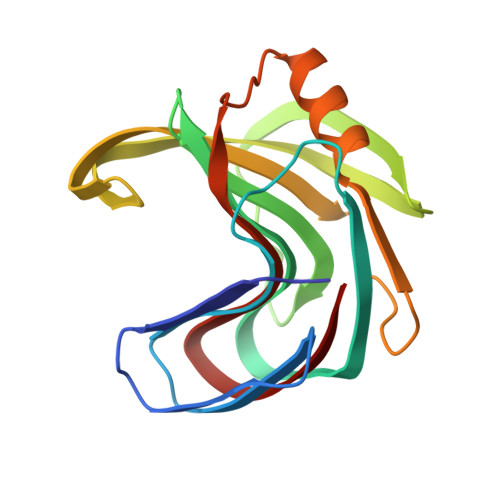
Identification of structural determinants for inhibition strength and specificity of wheat xylanase inhibitors TAXI-IA and TAXI-IIA.
Pollet, A., Sansen, S., Raedschelders, G., Gebruers, K., Rabijns, A., Delcour, J.A., Courtin, C.M.(2009) FEBS J 276: 3916-3927
- PubMed: 19769747 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2009.07105.x
- Primary Citation Related Structures:
2B42, 3HD8 - PubMed Abstract:
Triticum aestivum xylanase inhibitor (TAXI)-type inhibitors are active against microbial xylanases from glycoside hydrolase family 11, but the inhibition strength and the specificity towards different xylanases differ between TAXI isoforms. Mutational and biochemical analyses of TAXI-I, TAXI-IIA and Bacillus subtilis xylanase A showed that inhibition strength and specificity depend on the identity of only a few key residues of inhibitor and xylanase [Fierens K et al. (2005) FEBS J 272, 5872-5882; Raedschelders G et al. (2005) Biochem Biophys Res Commun335, 512-522; Sorensen JF & Sibbesen O (2006) Protein Eng Des Sel 19, 205-210; Bourgois TM et al. (2007) J Biotechnol 130, 95-105]. Crystallographic analysis of the structures of TAXI-IA and TAXI-IIA in complex with glycoside hydrolase family 11 B. subtilis xylanase A now provides a substantial explanation for these observations and a detailed insight into the structural determinants for inhibition strength and specificity. Structures of the xylanaseinhibitor complexes show that inhibition is established by loop interactions with active-site residues and substrate-mimicking contacts in the binding subsites. The interaction of residues Leu292 of TAXI-IA and Pro294 of TAXI-IIA with the -2 glycon subsite of the xylanase is shown to be critical for both inhibition strength and specificity. Also, detailed analysis of the interaction interfaces of the complexes illustrates that the inhibition strength of TAXI is related to the presence of an aspartate or asparagine residue adjacent to the acid/base catalyst of the xylanase, and therefore to the pH optimum of the xylanase. The lower the pH optimum of the xylanase, the stronger will be the interaction between enzyme and inhibitor, and the stronger the resulting inhibition.
- Laboratory of Food Chemistry and Biochemistry, Katholieke Universiteit Leuven, Belgium.
Organizational Affiliation: