Molecular characterization of a B-ring unsaturated estrogen: implications for conjugated equine estrogen components of premarin.
Hsieh, R.W., Rajan, S.S., Sharma, S.K., Greene, G.L.(2008) Steroids 73: 59-68
- PubMed: 17949766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.steroids.2007.08.014
- Primary Citation Related Structures:
2B1Z - PubMed Abstract:
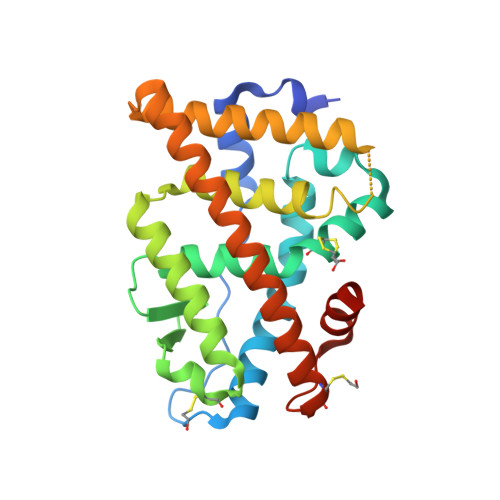
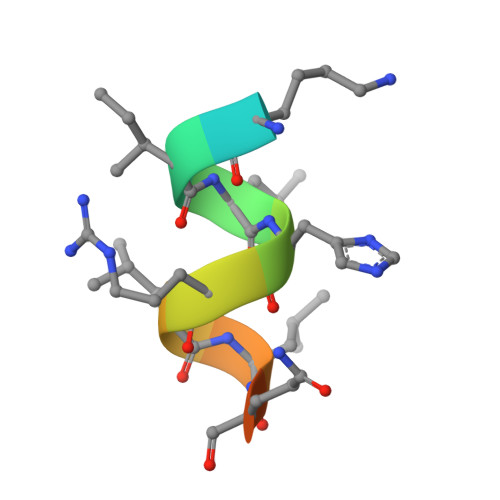
Conjugated equine estrogens (CEEs) are routinely used for hormone replacement therapy (HRT), making it important to understand the activities of individual estrogenic components. Although 17beta-estradiol (17beta-E2), the most potent estrogen in CEE, has been extensively characterized, the actions of nine additional less potent estrogens are not well understood. Structural differences between CEEs and 17beta-E2 result in altered interactions with the two estrogen receptors (ERalpha and ERbeta) and different biological activities. To better understand these interactions, we have determined the crystal structure of the CEE analog, 17beta-methyl-17alpha-dihydroequilenin (NCI 122), in complex with the ERalpha ligand-binding domain and a peptide from the glucocorticoid receptor-interacting protein 1 (GRIP1) coactivator. NCI 122 has chemical properties, including an unsaturated B-ring and 17alpha-hydroxyl group, which are shared with some of the estrogens found in CEEs. Structural analysis of the NCI 122-ERalpha LBD-GRIP1 complex, combined with biochemical and cell-based comparisons of CEE components, suggests that factors such as decreased ligand flexibility, decreased ligand hydrophobicity and loss of a hydrogen bond between the 17-hydroxyl group and His524, contribute significantly to the reduced potency of CEEs on ERalpha.
- The Ben May Department for Cancer Research, The University of Chicago, Chicago, IL 60637, USA.
Organizational Affiliation: