Design and synthesis of 6-anilinoindazoles as selective inhibitors of c-Jun N-terminal kinase-3
Swahn, B.M., Huerta, F., Kallin, E., Malmstrom, J., Weigelt, T., Viklund, J., Womack, P., Xue, Y., Ohberg, L.(2005) Bioorg Med Chem Lett 15: 5095-5099
- PubMed: 16140012 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2005.06.083
- Primary Citation Related Structures:
2B1P - PubMed Abstract:
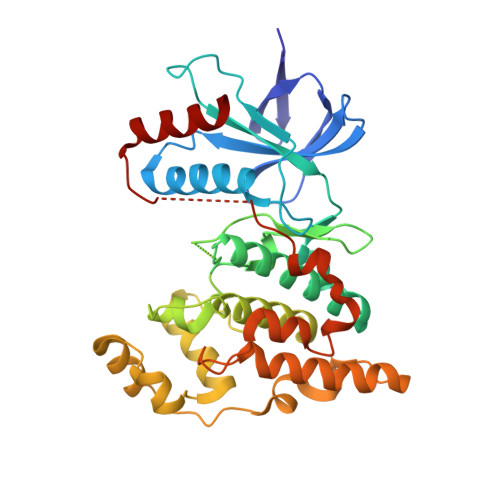
The structure-based design and synthesis of a new series of c-Jun N-terminal kinase-3 inhibitors with selectivity against JNK1 and p38alpha is reported. The novel series of substituted 6-anilinoindazoles were designed based on a combination of hits from high throughput screening and X-ray crystal structure information of the compounds crystallized into the JNK3 ATP binding active site.
- Department of Medicinal Chemistry, AstraZeneca R&D Södertälje, S-15185 Södertälje, Sweden. britt-marie.swahn@astrazeneca.com
Organizational Affiliation: