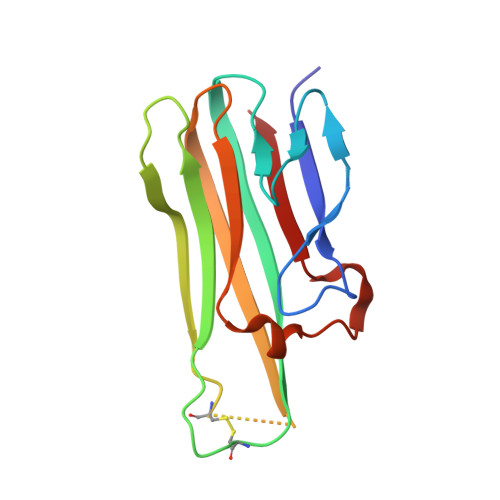
Small-molecule inhibition of TNF-alpha.
He, M.M., Smith, A.S., Oslob, J.D., Flanagan, W.M., Braisted, A.C., Whitty, A., Cancilla, M.T., Wang, J., Lugovskoy, A.A., Yoburn, J.C., Fung, A.D., Farrington, G., Eldredge, J.K., Day, E.S., Cruz, L.A., Cachero, T.G., Miller, S.K., Friedman, J.E., Choong, I.C., Cunningham, B.C.(2005) Science 310: 1022-1025
- PubMed: 16284179 Search on PubMed
- DOI: https://doi.org/10.1126/science.1116304
- Primary Citation Related Structures:
2AZ5 - PubMed Abstract:
We have identified a small-molecule inhibitor of tumor necrosis factor alpha (TNF-alpha) that promotes subunit disassembly of this trimeric cytokine family member. The compound inhibits TNF-alpha activity in biochemical and cell-based assays with median inhibitory concentrations of 22 and 4.6 micromolar, respectively. Formation of an intermediate complex between the compound and the intact trimer results in a 600-fold accelerated subunit dissociation rate that leads to trimer dissociation. A structure solved by x-ray crystallography reveals that a single compound molecule displaces a subunit of the trimer to form a complex with a dimer of TNF-alpha subunits.
- Sunesis Pharmaceuticals, Incorporated, 341 Oyster Point Boulevard, South San Francisco, CA 94080, USA.
Organizational Affiliation: