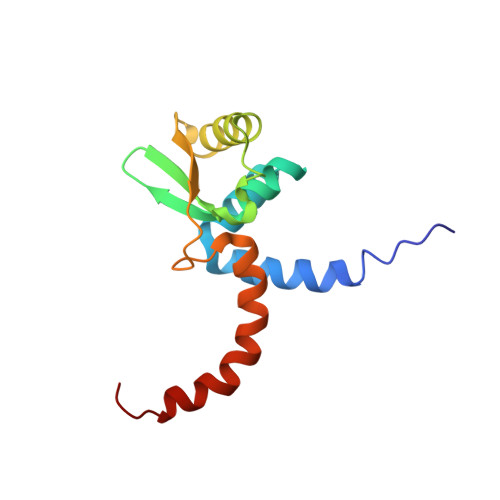
Solution structure of Domains IVa and V of the tau subunit of Escherichia coli DNA polymerase III and interaction with the alpha subunit.
Su, X.C., Jergic, S., Keniry, M.A., Dixon, N.E., Otting, G.(2007) Nucleic Acids Res 35: 2825-2832
- PubMed: 17452361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkm080
- Primary Citation Related Structures:
2AYA - PubMed Abstract:
The solution structure of the C-terminal Domain V of the tau subunit of E. coli DNA polymerase III was determined by nuclear magnetic resonance (NMR) spectroscopy. The fold is unique to tau subunits. Amino acid sequence conservation is pronounced for hydrophobic residues that form the structural core of the protein, indicating that the fold is representative for tau subunits from a wide range of different bacteria. The interaction between the polymerase subunits tau and alpha was studied by NMR experiments where alpha was incubated with full-length C-terminal domain (tau(C)16), and domains shortened at the C-terminus by 11 and 18 residues, respectively. The only interacting residues were found in the C-terminal 30-residue segment of tau, most of which is structurally disordered in free tau(C)16. Since the N- and C-termini of the structured core of tau(C)16 are located close to each other, this limits the possible distance between alpha and the pentameric deltatau2gammadelta' clamp-loader complex and, hence, between the two alpha subunits involved in leading- and lagging-strand DNA synthesis. Analysis of an N-terminally extended construct (tau(C)22) showed that tau(C)14 presents the only part of Domains IVa and V of tau which comprises a globular fold in the absence of other interaction partners.
- Research School of Chemistry, Australian National University, Canberra ACT 0200, Australia.
Organizational Affiliation: