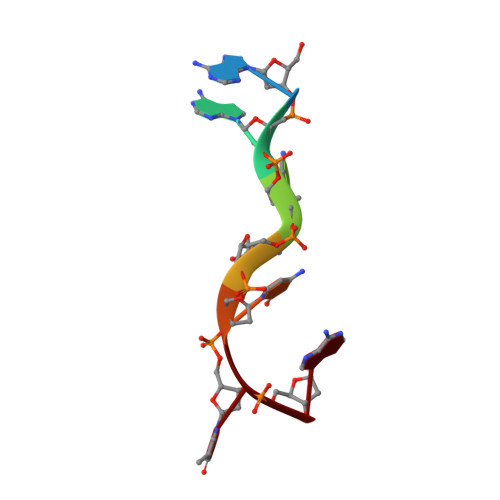
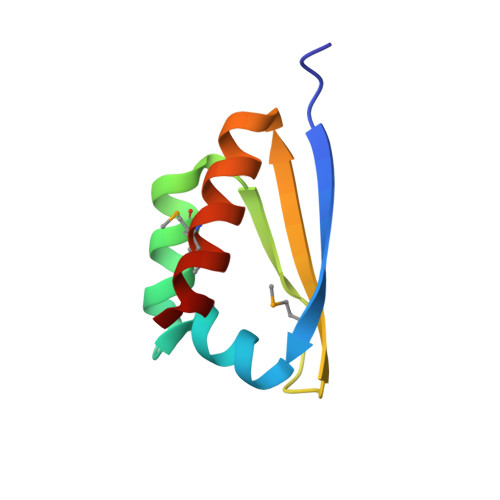
Crystal Structure of the First KH Domain of Human Poly(C)-binding Protein-2 in Complex with a C-rich Strand of Human Telomeric DNA at 1.7 A
Du, Z., Lee, J.K., Tjhen, R., Li, S., Pan, H., Stroud, R.M., James, T.L.(2005) J Biological Chem 280: 38823-38830
- PubMed: 16186123 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M508183200
- Primary Citation Related Structures:
2AXY - PubMed Abstract:
Recognition of poly(C) DNA and RNA sequences in mammalian cells is achieved by a subfamily of the KH (hnRNP K homology) domain-containing proteins known as poly(C)-binding proteins (PCBPs). To reveal the molecular basis of poly(C) sequence recognition, we have determined the crystal structure, at 1.7-A resolution, of PCBP2 KH1 in complex with a 7-nucleotide DNA sequence (5'-AACCCTA-3') corresponding to one repeat of the human C-rich strand telomeric DNA. The protein-DNA interaction is mediated by the combination of several stabilizing forces including hydrogen bonding, electrostatic interactions, van der Waals contacts, and shape complementarities. Specific recognition of the three cytosine residues is realized by a dense network of hydrogen bonds involving the side chains of two conserved lysines and one glutamic acid. The co-crystal structure also reveals a protein-protein dimerization interface of PCBP2 KH1 located on the opposite side of the protein from the DNA binding groove. Numerous stabilizing protein-protein interactions, including hydrophobic contacts, stacking of aromatic side chains, and a large number of hydrogen bonds, indicate that the protein-protein interaction interface is most likely genuine. Interaction of PCBP2 KH1 with the C-rich strand of human telomeric DNA suggests that PCBPs may participate in mechanisms involved in the regulation of telomere/telomerase functions.
- Department of Pharmaceutical Chemistry, University of California, San Francisco, California 94143-2280, USA.
Organizational Affiliation: