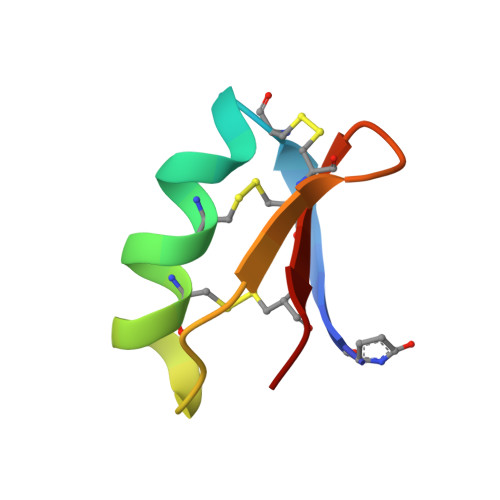
Solution structure of discrepin, a new K+-channel blocking peptide from the alpha-KTx15 subfamily.
Prochnicka-Chalufour, A., Corzo, G., Satake, H., Martin-Eauclaire, M.-F., Murgia, A.R., Prestipino, G., D'Suze, G., Possani, L.D., Delepierre, M.(2006) Biochemistry 45: 1795-1804
- PubMed: 16460026 Search on PubMed
- DOI: https://doi.org/10.1021/bi0519248
- Primary Citation Related Structures:
2AXK - PubMed Abstract:
Discrepin, isolated from the venom of the Venezuelan scorpion Tityus discrepans, blocks preferentially the I(A) currents of the voltage-dependent K+ channel of rat cerebellum granular cells in an irreversible way. It contains 38 amino acid residues with a pyroglutamic acid as the N-terminal residue [D'Suze, G., Batista, C. V., Frau, A., Murgia, A. R., Zamudio, F. Z., Sevcik, C., Possani, L. D., and Prestipino, G. (2004) Arch. Biochem. Biophys. 430, 256-63]. It is the most distinctive member of the alpha-KTx15 subfamily of scorpion toxins. Six members of the alpha-KTx15 subfamily have been reported so far to be specific for this subtype of the K+ channel; however, none of them have had their three-dimensional structure determined, and no information for the residues possibly involved in channel recognition and binding is available. Natural discrepin (n-discrepin) was prepared from scorpion venom, and its synthetic analogue (s-discrepin) was obtained by solid-phase synthesis. Analysis of two-dimensional 1H NMR spectra of n- and s-discrepin indicates that both peptides have the same structure. Here we report the solution structure of s-discrepin determined by NMR using 565 meaningful distance constraints derived from the volume integration of the two-dimensional NOESY spectrum, 22 dihedrals, and three hydrogen bonds. Discrepin displays the alpha/beta scaffold, characteristic of scorpion toxins. Some features of the proposed interacting surface between the toxin and channel as well as the opposite "alpha-helix surface" are discussed in comparison with those of other alpha-KTx15 members. Both n- and s-discrepin exhibit similar physiological actions as verified by patch-clamp and binding and displacement experiments.
- Unité de RMN des Biomolécules URA 2185 CNRS, Institut Pasteur, 28 Rue du Dr Roux, 75724 Paris Cedex 15, France.
Organizational Affiliation: