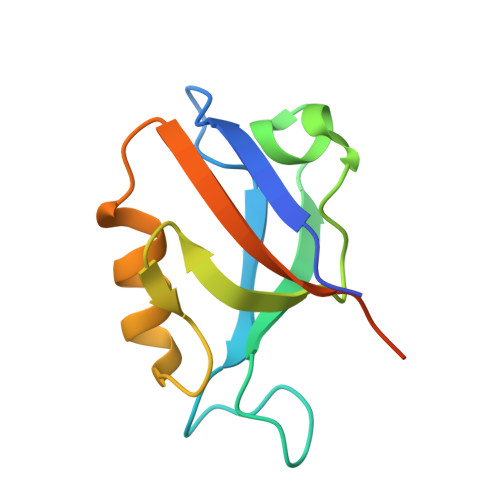
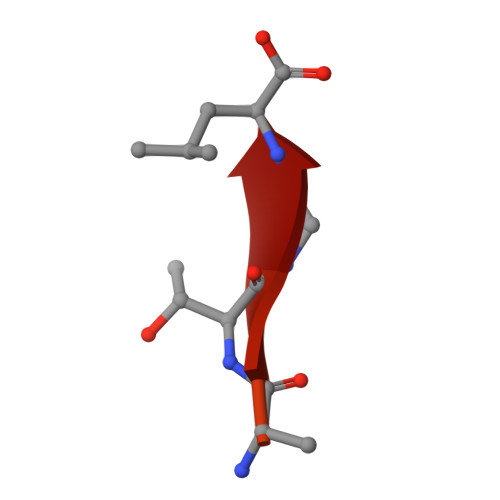
Crystal structure of the second PDZ domain of SAP97 in complex with a GluR-A C-terminal peptide
von Ossowski, I., Oksanen, E., von Ossowski, L., Cai, C., Sundberg, M., Goldman, A., Keinanen, K.(2006) FEBS J 273: 5219-5229
- PubMed: 17069616 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05521.x
- Primary Citation Related Structures:
2AWU, 2AWW, 2AWX, 2G2L - PubMed Abstract:
Synaptic targeting of GluR-A subunit-containing glutamate receptors involves an interaction with synapse-associated protein 97 (SAP97). The C-terminus of GluR-A, which contains a class I PDZ ligand motif (-x-Ser/Thr-x-phi-COOH where phi is an aliphatic amino acid) associates preferentially with the second PDZ domain of SAP97 (SAP97(PDZ2)). To understand the structural basis of this interaction, we have determined the crystal structures of wild-type and a SAP97(PDZ2) variant in complex with an 18-mer C-terminal peptide (residues 890-907) of GluR-A and of two variant PDZ2 domains in unliganded state at 1.8-2.44 A resolutions. SAP97(PDZ2) folds to a compact globular domain comprising six beta-strands and two alpha-helices, a typical architecture for PDZ domains. In the structure of the peptide complex, only the last four C-terminal residues of the GluR-A are visible, and align as an antiparallel beta-strand in the binding groove of SAP97(PDZ2). The free carboxylate group and the aliphatic side chain of the C-terminal leucine (Leu907), and the hydroxyl group of Thr905 of the GluR-A peptide are engaged in essential class I PDZ interactions. Comparison between the free and complexed structures reveals conformational changes which take place upon peptide binding. The betaAlpha-betaBeta loop moves away from the C-terminal end of alphaB leading to a slight opening of the binding groove, which may better accommodate the peptide ligand. The two conformational states are stabilized by alternative hydrogen bond and coulombic interactions of Lys324 in betaAlpha-betaBeta loop with Asp396 or Thr394 in betaBeta. Results of in vitro binding and immunoprecipitation experiments using a PDZ motif-destroying L907A mutation as well as the insertion of an extra alanine residue between the C-terminal Leu907 and the stop codon are also consistent with a 'classical' type I PDZ interaction between SAP97 and GluR-A C-terminus.
- Department of Biological and Environmental Sciences (Division of Biochemistry), University of Helsinki, Finland.
Organizational Affiliation: