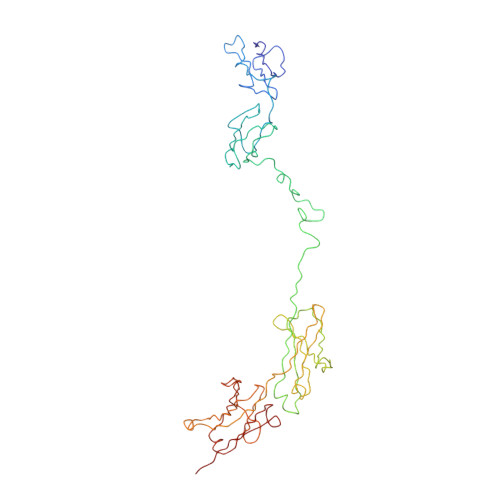
Extended Flexible Linker Structures in the Complement Chimaeric Conjugate CR2-Ig by Scattering, Analytical Ultracentrifugation and Constrained Modelling: Implications for Function and Therapy.
Gilbert, H.E., Aslam, M., Guthridge, J.M., Holers, V.M., Perkins, S.J.(2006) J Mol Biology 356: 397-412
- PubMed: 16375923 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.050
- Primary Citation Related Structures:
2ATY - PubMed Abstract:
Complement receptor 2 (CR2; CD21) is a membrane-bound regulator of complement activation, being comprised of 15 or 16 short complement repeat (SCR) domains. A recombinant glycosylated human CR2 SCR 1-2 domain pair was engineered with the Fc fragment of a mouse IgG1 antibody to create a chimaera CR2-Ig containing the major ligand binding domains. Such a chimaera has therapeutic potential as a complement inhibitor or immune modulator. X-ray and neutron scattering and analytical ultracentrifugation identified its domain structure in solution, and provided a comparison with controversial folded-back crystal structures for deglycosylated CR2 SCR 1-2. The radius of gyration R(G) of CR2-Ig was determined to be 5.39(+/-0.14) nm and 5.29(+/-0.01) nm by X-ray and neutron scattering, respectively. The maximum dimension of CR2-Ig was determined to be 17 nm. The molecular mass of CR2-Ig ranged between 101,000 Da and 107,000 Da as determined by neutron scattering and sedimentation equilibrium, in good agreement with the sequence-derived value of 106,600 Da. Sedimentation velocity gave a sedimentation coefficient of 4.49(+/-0.11) S. Stereochemically complete models for CR2-Ig were constructed from crystal structures for the CR2 SCR 1-2 and mouse IgG1 Fc fragments. The two SCR domains and the Fc fragment were joined by randomised conformational peptides. The analysis of 35,000 possible CR2-Ig models showed that only those models in which the two SCR domains were arranged in an open V-shape in random orientations about the Fc fragment accounted for the scattering and sedimentation data. It was not possible to define one single conformational family of Fab-like fragment relative to the Fc fragment. This flexibility is attributed to the relatively long linker sequence and the absence of the antibody light chain from CR2-Ig. The modelling also confirmed that the structure of CR2 SCR 1-2 is more extended in solution than in its crystal structure.
- Department of Biochemistry and Molecular Biology, Darwin Building, University College London, UK.
Organizational Affiliation: