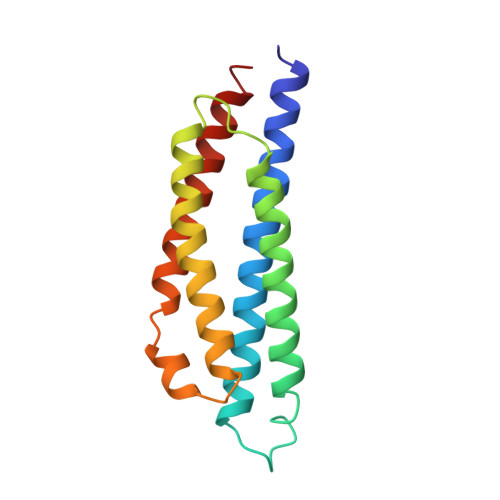
The three-dimensional structure of the aspartate receptor from Escherichia coli.
Bowie, J.U., Pakula, A.A., Simon, M.I.(1995) Acta Crystallogr D Biol Crystallogr 51: 145-154
- PubMed: 15299315
- DOI: https://doi.org/10.1107/S0907444994010498
- Primary Citation Related Structures:
2ASR - PubMed Abstract:
The crystal structure of the periplasmic domain of the aspartate receptor from Escherichia coli has been solved and refined to an R-factor of 0.203 at 2.3 A, resolution. The dimeric protein is largely helical, with four helices from each monomer forming a four-helix bundle. The dimer interface is constructed from four helices, two from each subunit, also packed together in a four-helix bundle arrangement. A sulfate ion occupies the aspartate-binding site. All hydrogen bonds made to aspartate are substituted by direct or water-mediated hydrogen bonds to the sulfate. Comparison of the Escherichia coli aspartate-receptor structure with that of Salmonella typhimurium [Milburn, Prive, Milligan, Scott, Yeh, Jancarik, Koshland & Kim (1991). Science, 254, 1342-1347; Scott, Milligan, Milburn, Prive, Yeh, Koshland & Kim (1993). J. Mol. Biol. 232, 555-573] reveals strong conservation in the structure of the monomer, but more divergence in the orientation of the subunits with respect to one another. Mutations that render the Escherichia coli receptor incapable of responding to maltose are either located in spatially conserved sites or in regions of the structures that have high temperature factors and are therefore likely to be quite flexible. The inability of the receptor from Salmonella typhimurium to respond to maltose may, therefore, be because of differences in amino acids located on the binding surface rather than structural differences.
- Department of Chemistry and Biochemistry, and DOE Laboratory of Structural Biology and Molecular Medicine, University of California, Los Angeles 90024, USA.
Organizational Affiliation: