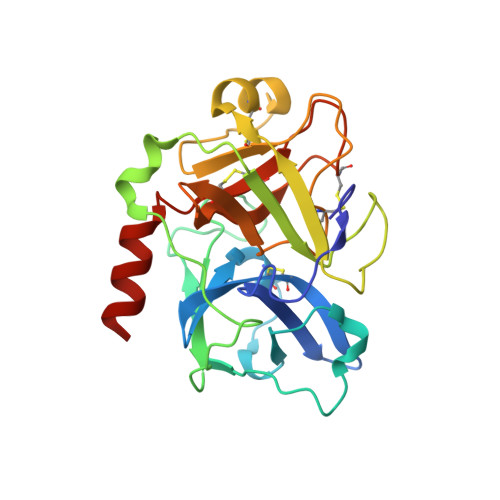
Expression, crystallization, and three-dimensional structure of the catalytic domain of human plasma kallikrein.
Tang, J., Yu, C.L., Williams, S.R., Springman, E., Jeffery, D., Sprengeler, P.A., Estevez, A., Sampang, J., Shrader, W., Spencer, J., Young, W., McGrath, M., Katz, B.A.(2005) J Biological Chem 280: 41077-41089
- PubMed: 16199530 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M506766200
- Primary Citation Related Structures:
2ANW, 2ANY - PubMed Abstract:
Plasma kallikrein is a serine protease that has many important functions, including modulation of blood pressure, complement activation, and mediation and maintenance of inflammatory responses. Although plasma kallikrein has been purified for 40 years, its structure has not been elucidated. In this report, we described two systems (Pichia pastoris and baculovirus/Sf9 cells) for expression of the protease domain of plasma kallikrein, along with the purification and high resolution crystal structures of the two recombinant forms. In the Pichia pastoris system, the protease domain was expressed as a heterogeneously glycosylated zymogen that was activated by limited trypsin digestion and treated with endoglycosidase H deglycosidase to reduce heterogeneity from the glycosylation. The resulting protein was chromatographically resolved into four components, one of which was crystallized. In the baculovirus/Sf9 system, homogeneous, crystallizable, and nonglycosylated protein was expressed after mutagenizing three asparagines (the glycosylation sites) to glutamates. When assayed against the peptide substrates, pefachrome-PK and oxidized insulin B chain, both forms of the protease domain were found to have catalytic activity similar to that of the full-length protein. Crystallization and x-ray crystal structure determination of both forms have yielded the first three-dimensional views of the catalytic domain of plasma kallikrein. The structures, determined at 1.85 A for the endoglycosidase H-deglycosylated protease domain produced from P. pastoris and at 1.40 A for the mutagenically deglycosylated form produced from Sf9 cells, show that the protease domain adopts a typical chymotrypsin-like serine protease conformation. The structural information provides insights into the biochemical and enzymatic properties of plasma kallikrein and paves the way for structure-based design of protease inhibitors that are selective either for or against plasma kallikrein.
- Department of Structural Chemistry, Celera Genomics, South San Francisco, California 94080, USA.
Organizational Affiliation: