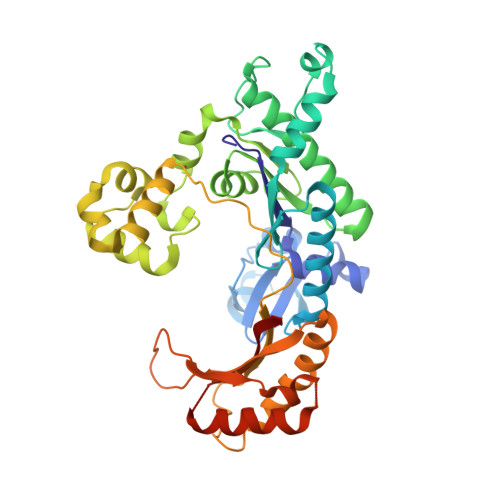
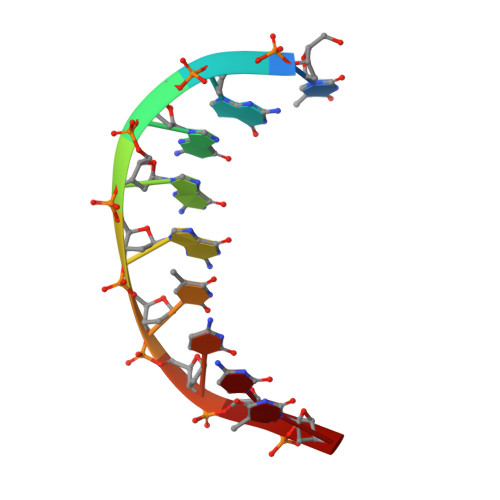
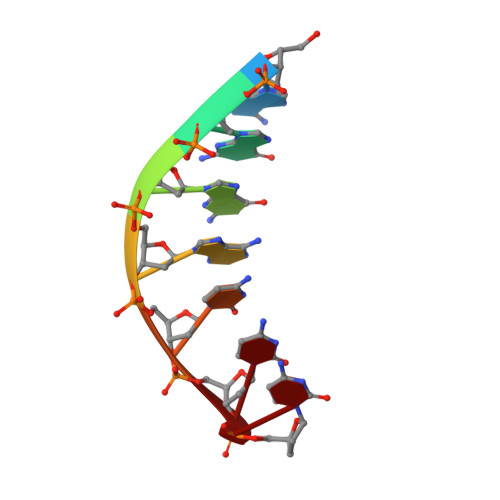
Human DNA Polymerase iota Incorporates dCTP Opposite Template G via a G.C+ Hoogsteen Base Pair.
Nair, D.T., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2005) Structure 13: 1569-1577
- PubMed: 16216587 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.08.010
- Primary Citation Related Structures:
2ALZ - PubMed Abstract:
Human DNA polymerase iota (hPoliota), a member of the Y family of DNA polymerases, differs in remarkable ways from other DNA polymerases, incorporating correct nucleotides opposite template purines with a much higher efficiency and fidelity than opposite template pyrimidines. We present here the crystal structure of hPoliota bound to template G and incoming dCTP, which reveals a G.C + Hoogsteen base pair in a DNA polymerase active site. We show that the hPoliota active site has evolved to favor Hoogsteen base pairing, wherein the template sugar is fixed in a cavity that reduces the C1'-C1' distance across the nascent base pair from approximately 10.5 A in other DNA polymerases to 8.6 A in hPoliota. The rotation of G from anti to syn is then largely in response to this curtailed C1'-C1' distance. A G.C+ Hoogsteen base pair suggests a specific mechanism for hPoliota's ability to bypass N(2)-adducted guanines that obstruct replication.
- Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, Box 1677, 1425 Madison Avenue, New York, New York 10029, USA.
Organizational Affiliation: