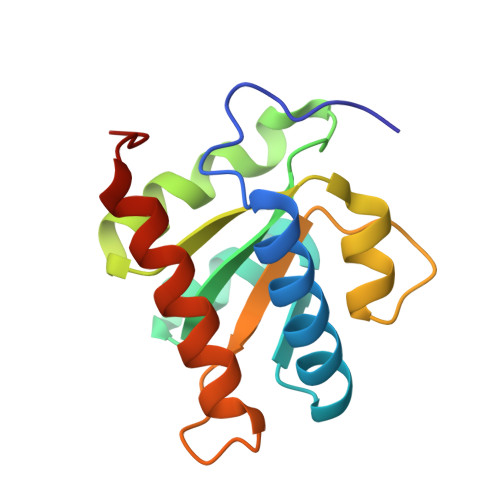
Analysis of pre-mRNA and pre-rRNA processing factor Snu13p structure and mutants.
Dobbyn, H.C., McEwan, P.A., Krause, A., Novak-Frazer, L., Bella, J., O'Keefe, R.T.(2007) Biochem Biophys Res Commun 360: 857-862
- PubMed: 17631273 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.06.163
- Primary Citation Related Structures:
2ALE - PubMed Abstract:
Snu13p is a Saccharomyces cerevisiae protein essential for pre-messenger RNA splicing and pre-ribosomal RNA processing. Snu13p binds U4 snRNA of the spliceosome and box C/D snoRNAs of the pre-ribosomal RNA processing machinery to induce assembly of each ribonucleoprotein complex. Here, we present structural and biochemical analysis of Snu13p. The crystal structure of Snu13p reveals a region of the protein which could be important for protein interaction during ribonucleoprotein assembly. Using the structure of Snu13p we have designed the first temperature-sensitive mutants in Snu13p, L67W and I102A. Wild-type and mutant Snu13p proteins were assayed for binding to U4 snRNA and U3 snoRNA. Both temperature-sensitive mutants displayed significantly reduced RNA binding compared to wild-type protein. As the temperature-sensitive mutations are not in the known RNA binding region of Snu13p this indicates that these mutants indirectly influence the RNA binding properties of Snu13p. This work provides insight into Snu13p function during ribonucleoprotein assembly.
- School of Pharmacy, Centre for Biomolecular Sciences, University of Nottingham, Nottingham NG7 2RD, UK.
Organizational Affiliation: