Structure of replicative DNA polymerase provides insigts into the mechanisms for processivity, frameshifting and editing
Brieba, L., Ellenberger, T.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| T7 DNA polymerase | E [auth A], G [auth F] | 704 | Escherichia phage T7 | Mutation(s): 2 Gene Names: 5 EC: 2.7.7.7 (PDB Primary Data), 3.1.11 (UniProt) | 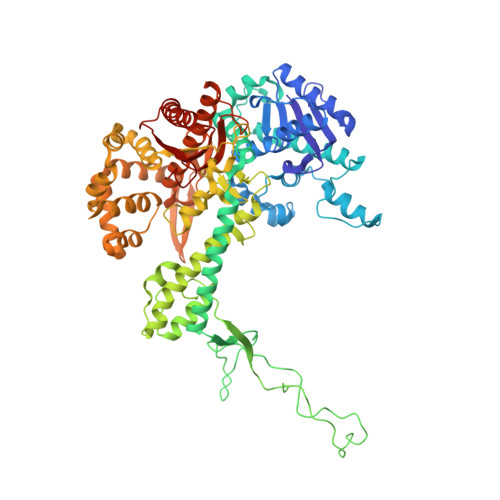 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00581 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| thioredoxin 1 | F [auth B], H [auth I] | 108 | Escherichia coli | Mutation(s): 0 Gene Names: trxA, fipA, tsnC | 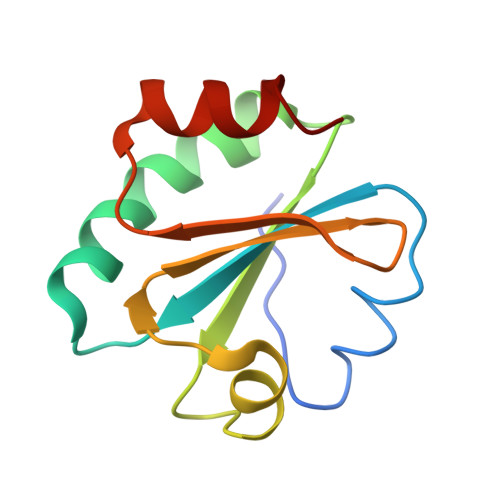 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0AA25 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA Primer | A [auth P], C [auth X] | 22 | N/A | 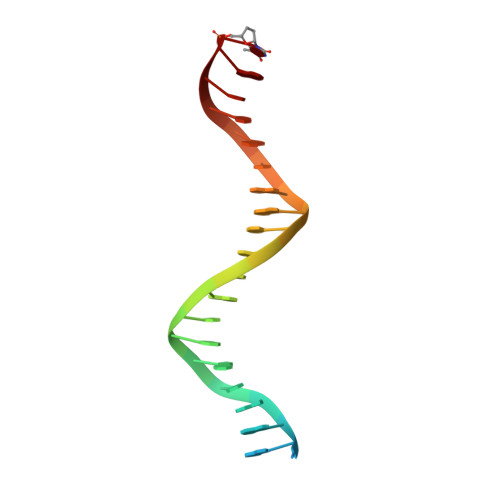 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA Template | B [auth T], D [auth Z] | 26 | N/A | 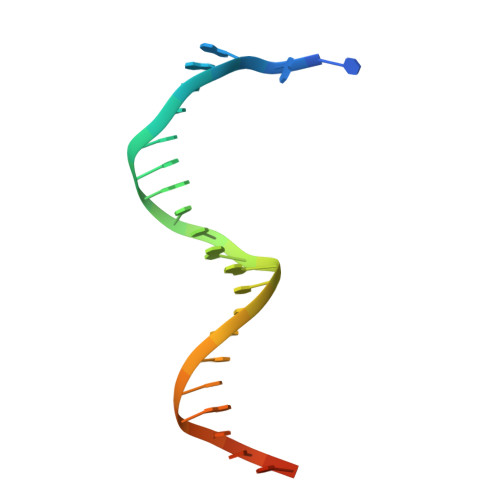 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 168.312 | α = 90 |
| b = 169.235 | β = 90 |
| c = 179.788 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data collection |
| TRUNCATE | data reduction |
| CNS | refinement |
| HKL-2000 | data reduction |
| CCP4 | data scaling |
| CNS | phasing |